 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g70270.2 | ||||||||
| Common Name | ARF2, ARF4, LOC4326957, Os01g0927600, OSJNBa0093F16.35 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 809aa MW: 90854.7 Da PI: 6.4775 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 69.6 | 4.1e-22 | 129 | 230 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldg 86
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d+++ W++++i+r++++r++l++GW+ Fv++++L +gD ++F +
LOC_Os01g70270.2 129 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHSMDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--R 217
99*****************************7555544..58************************************************..4 PP
SSEE..EEEEE-S CS
B3 87 rsefelvvkvfrk 99
+++el+v+v+r+
LOC_Os01g70270.2 218 GENGELRVGVRRA 230
49999*****996 PP
| |||||||
| 2 | Auxin_resp | 113.6 | 1.9e-37 | 255 | 336 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++svGmRf+m+fe+e+++e+r++Gt++g ++ldp Wp+S WrsLk
LOC_Os01g70270.2 255 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFEGEEAPEQRFTGTIIGSENLDP-VWPESSWRSLK 336
89*********************************************************************.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 8.63E-40 | 121 | 258 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.3E-39 | 122 | 244 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.36E-19 | 127 | 229 | No hit | No description |
| PROSITE profile | PS50863 | 11.265 | 129 | 231 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.5E-21 | 129 | 231 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.1E-20 | 129 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.9E-34 | 255 | 336 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 8.5E-7 | 673 | 728 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 4.51E-13 | 688 | 769 | No hit | No description |
| PROSITE profile | PS51745 | 26.238 | 692 | 785 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 9.8E-4 | 710 | 765 | No hit | No description |
| Pfam | PF02309 | 2.6E-8 | 734 | 777 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009049 | anatomy | inflorescence | ||||
| PO:0001083 | developmental stage | inflorescence development stage | ||||
| PO:0007006 | developmental stage | IL.00 inflorescence just visible stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 809 aa Download sequence Send to blast |
MPPAAMAPPP PPQGSSTGDP LYDELWHACA GPLVTVPRVG DLVFYFPQGH IEQVEASMNQ 60 VADSQMRLYD LPSKLLCRVL NVELKAEQDT DEVYAQVMLM PEPEQNEMAV EKTTPTSGPV 120 QARPPVRSFC KTLTASDTST HGGFSVLRRH ADECLPPLDM TQSPPTQELV AKDLHSMDWR 180 FRHIFRGQPR RHLLQSGWSV FVSSKRLVAG DAFIFLRGEN GELRVGVRRA MRQLSNVPSS 240 VISSQSMHLG VLATAWHAIN TKSMFTVYYK PRTSPSEFII PYDQYMESVK NNYSVGMRFR 300 MRFEGEEAPE QRFTGTIIGS ENLDPVWPES SWRSLKVRWD EPSTIPRPDR VSPWKIEPAS 360 SPPVNPLPLS RVKRPRPNAP PASPESPILT KEAATKVDTD PAQAQRSQNS TVLQGQEQMT 420 LRSNLTESND SDVTAHKPMM WSPSPNAAKA HPLTFQQRPP MDNWMQLGRR ETDFKDVRSG 480 SQSFGDSPGF FMQNFDEAPN RLTSFKNQFQ DQGSARHFSD PYYYVSPQPS LTVESSTQMH 540 TDSKELHFWN GQSTVYGNSR DRPQNFRFEQ NSSSWLNQSF ARPEQPRVIR PHASIAPVEL 600 EKTEGSGFKI FGFKVDTTNA PNNHLSSPMA ATHEPMLQTP SSLNQLQPVQ TDCIPEVSVS 660 TAGTATENEK SGQQAQQSSK DVQSKTQVAS TRSCTKVHKQ GVALGRSVDL SKFSNYDELK 720 AELDKMFEFD GELVSSNKNW QIVYTDNEGD MMLVGDDPWE EFCSIVRKIY IYTKEEVQKM 780 NSKSNAPRKD DSSENEKGHL PMPNKSDN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-169 | 14 | 361 | 12 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-169 | 14 | 361 | 12 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-169 | 14 | 361 | 12 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-169 | 14 | 361 | 12 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-169 | 14 | 361 | 12 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 372 | 376 | KRPRP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.26144 | 0.0 | callus| flower| leaf| panicle| root| seed| stem| vegetative meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32982332 | 0.0 | ||||
| Expression Atlas | Q5JK20 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
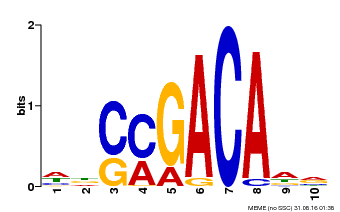
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g70270.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g70270 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072309 | 0.0 | AK072309.1 Oryza sativa Japonica Group cDNA clone:J023022C23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015650953.1 | 0.0 | auxin response factor 4 | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A0E0N733 | 0.0 | A0A0E0N733_ORYRU; Auxin response factor | ||||
| STRING | OS01T0927600-01 | 0.0 | (Oryza sativa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g70270.2 |
| Entrez Gene | 4326957 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



