 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g60270.2 | ||||||||
| Common Name | LOC4327441, Os01g0818400, OsJ_03880, P0454H12.31, WOX8 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | WOX | ||||||||
| Protein Properties | Length: 268aa MW: 30238.7 Da PI: 6.3521 | ||||||||
| Description | WOX family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 67.6 | 1.6e-21 | 90 | 150 | 2 | 57 |
T--SS--HHHHHHHHHHHHH.SSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 2 rkRttftkeqleeLeelFek.nrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
r+R+t+t+ ql++Le+ F + n +ps++++++++++l +++e++V++WFqNrRa+ k+
LOC_Os01g60270.2 90 RQRWTPTPMQLQILENIFDQgNGTPSKQKIKDITAELsqhgQISETNVYNWFQNRRARSKR 150
89*****************99**************************************97 PP
| |||||||
| 2 | Wus_type_Homeobox | 104.3 | 8.2e-34 | 89 | 152 | 2 | 65 |
Wus_type_Homeobox 2 artRWtPtpeQikiLeelyksGlrtPnkeeiqritaeLeeyGkiedkNVfyWFQNrkaRerqkq 65
ar+RWtPtp Q++iLe+++++G tP+k++i++itaeL+++G+i+++NV++WFQNr+aR+++kq
LOC_Os01g60270.2 89 ARQRWTPTPMQLQILENIFDQGNGTPSKQKIKDITAELSQHGQISETNVYNWFQNRRARSKRKQ 152
789***********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-17 | 85 | 155 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 14.835 | 86 | 151 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.9E-14 | 88 | 155 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.01E-18 | 89 | 153 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.45E-13 | 90 | 152 | No hit | No description |
| Pfam | PF00046 | 2.6E-19 | 90 | 150 | IPR001356 | Homeobox domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007275 | Biological Process | multicellular organism development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0007130 | developmental stage | sporophyte reproductive stage | ||||
| PO:0007606 | developmental stage | gynoecium development stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MEWDKAKASS GEAVDDRGGG EGGLGYVKVM TDEQMEVLRK QISIYATICE QLVEMHRALT 60 AQQDSIAGMR LGNLYCDPLM VPGGHKITAR QRWTPTPMQL QILENIFDQG NGTPSKQKIK 120 DITAELSQHG QISETNVYNW FQNRRARSKR KQAALPNNNA ESEAEADEES PTDKKPKSDR 180 PLHQNIAMRD HNSERISEMH HFDTEHEQIR RMMYASNDSS SRSSGSLGQM SFYDNVMSNP 240 RIDHFLGKVE SPGSFPHMRS GESFDMY* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 143 | 151 | RRARSKRKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.24893 | 0.0 | callus| flower| leaf| panicle| root| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 116639018 | 0.0 | ||||
| Expression Atlas | Q5QMM3 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor which may be involved in developmental processes. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00669 | PBM | Transfer from GRMZM2G038252 | Download |
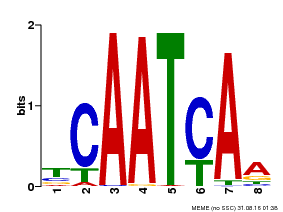
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g60270.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g60270 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK068585 | 0.0 | AK068585.1 Oryza sativa Japonica Group cDNA clone:J013151P17, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015626455.1 | 0.0 | WUSCHEL-related homeobox 8 | ||||
| Swissprot | Q5QMM3 | 0.0 | WOX8_ORYSJ; WUSCHEL-related homeobox 8 | ||||
| TrEMBL | A0A0E0FUZ4 | 0.0 | A0A0E0FUZ4_ORYNI; Uncharacterized protein | ||||
| TrEMBL | B8ABA9 | 0.0 | B8ABA9_ORYSI; Uncharacterized protein | ||||
| STRING | OS01T0818400-01 | 0.0 | (Oryza sativa) | ||||
| STRING | ONIVA01G40180.2 | 0.0 | (Oryza nivara) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G35550.1 | 2e-71 | WUSCHEL related homeobox 13 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g60270.2 |
| Entrez Gene | 4327441 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



