 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g54210.1 | ||||||||
| Common Name | LOC4326175, Os01g0745700, OSJNBa0014K08.27-1, OSNPB_010745700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 388aa MW: 40610.5 Da PI: 6.6245 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 55.2 | 9.3e-18 | 264 | 298 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C t kTp+WR gp g+ktLCnaCG++y++ +l
LOC_Os01g54210.1 264 CLHCETDKTPQWRTGPMGPKTLCNACGVRYKSGRL 298
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF016992 | 6.7E-71 | 15 | 347 | IPR016679 | Transcription factor, GATA, plant |
| PROSITE profile | PS50114 | 12.453 | 258 | 294 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.3E-18 | 258 | 308 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 4.4E-15 | 258 | 295 | IPR013088 | Zinc finger, NHR/GATA-type |
| SuperFamily | SSF57716 | 1.24E-15 | 260 | 321 | No hit | No description |
| CDD | cd00202 | 1.58E-14 | 263 | 314 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 264 | 289 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 1.4E-15 | 264 | 298 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009066 | anatomy | anther | ||||
| PO:0001004 | developmental stage | anther development stage | ||||
| PO:0007130 | developmental stage | sporophyte reproductive stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 388 aa Download sequence Send to blast |
MEVTAEFGGA YYGGAAGREK KALQQGCGDH FAVDDLLVLP YGEEDETTRE GEATGGKEEA 60 AGFGNASADS STITALDSCS NSFGLADGDF PGELCEPYDQ LAELEWLSNY MNEGDDAFAT 120 EDLQKLQLIS GIPSGGFSTA SVPSAQAQAA SAAASMAVQP GGFLPEAPVP AKARSKRSRA 180 APGNWSSRLL VLPPPPASPP SPASMAISPA ESGVSAHAFP IKKPSKPAKK KDAPAPPAQA 240 QLSSVPVHSG GSAPAAAAGE GRRCLHCETD KTPQWRTGPM GPKTLCNACG VRYKSGRLVP 300 EYRPAASPTF MVSKHSNSHR KVLELRRQKE MHQQTPHHHQ PQVAAAGGVG SLMHMQSSML 360 FDGVSPVVSG DDFLIHHHLR TDFRPPI* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.27911 | 0.0 | callus| flower| leaf| panicle| root| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 37991519 | 0.0 | ||||
| Expression Atlas | Q8LIZ3 | - | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
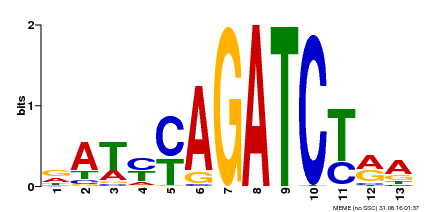
| Motif ID | Method | Source | Motif file |
| MP00530 | DAP | Transfer from AT5G25830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g54210.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g54210 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK121896 | 0.0 | AK121896.1 Oryza sativa Japonica Group cDNA clone:J033105E11, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631175.1 | 0.0 | GATA transcription factor 12 | ||||
| TrEMBL | Q8LIZ3 | 0.0 | Q8LIZ3_ORYSJ; GATA transcription factor | ||||
| STRING | OS01T0745700-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3025 | 31 | 78 | Representative plant | OGRP68 | 17 | 287 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G25830.1 | 6e-49 | GATA transcription factor 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g54210.1 |
| Entrez Gene | 4326175 |



