 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g48130.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 336aa MW: 36302.7 Da PI: 9.0816 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 105 | 9.8e-33 | 71 | 211 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkk......leleev.ikev.diykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratk...... 79
lp+G++F+Ptd+el+ e+L+ k++ + ++++ i+ +i++++P++Lp v ++ +fF++ +k+y+tg+rk+r+++
LOC_Os01g48130.2 71 LPAGVKFDPTDQELL-EHLEGKARPDArklhplIDEFIPtIEGEnGICYTHPERLP-GVGKDGLIRHFFHRPSKAYTTGTRKRRKVHtdeqgg 161
799************.99****998885665443333333665557**********.555555677******************744456555 PP
NAM 80 sgyWkatgkdkevlskkgelvglkktLvfy..kgrapkgektdWvmheyrl 128
+ +W++tgk+++v++ +g+l+g kk+Lv+y g+++k+ekt+Wvmh+y+l
LOC_Os01g48130.2 162 ETRWHKTGKTRPVFT-GGKLKGYKKILVLYtnYGKQRKPEKTNWVMHQYHL 211
778************.9*************6657999999*********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.19E-37 | 63 | 230 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 33.334 | 71 | 230 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.6E-15 | 72 | 211 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MTWCNSFNDV RAVENNLATA AAVAAAKKQQ QQQQVSQHVN LIKTCPSCGH RAQYEQLQAA 60 AAATIQDLPG LPAGVKFDPT DQELLEHLEG KARPDARKLH PLIDEFIPTI EGENGICYTH 120 PERLPGVGKD GLIRHFFHRP SKAYTTGTRK RRKVHTDEQG GETRWHKTGK TRPVFTGGKL 180 KGYKKILVLY TNYGKQRKPE KTNWVMHQYH LGSDEEEKDG ELVVSKVFYQ TQPRQCGGGS 240 AATAKDLSVD LVAGNNIKAS NAAAEHHHND GVGGGGHGGN NSSMLKEAAG IVDFYNPAAA 300 LIGYSQAAPN NRAAASAHLT MPNFEVHTGG AGFGP* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.9851 | 0.0 | callus| panicle| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32988003 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in protoxylem and elongating interfascicular fiber cells of elongating internodes, developing metaxylem cells and interfascicular fibers of non-elongating internodes and developing secondary xylem of roots. {ECO:0000269|PubMed:18952777}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that plays a regulatory role in the development of secondary cell wall fibers (PubMed:18952777,PubMed:22133261). Involved in the regulation of cellulose and hemicellulose biosynthetic genes as well as of genes involved in lignin polymerization and signaling (PubMed:22133261). Is not a direct target of SND1 (PubMed:18952777). {ECO:0000269|PubMed:18952777, ECO:0000269|PubMed:22133261}. | |||||
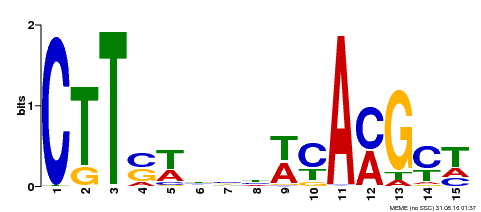
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00459 | DAP | Transfer from AT4G28500 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g48130.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g48130 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK102794 | 0.0 | AK102794.1 Oryza sativa Japonica Group cDNA clone:J033108B03, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015626614.1 | 0.0 | NAC domain-containing protein 73 isoform X2 | ||||
| Swissprot | O49459 | 1e-126 | NAC73_ARATH; NAC domain-containing protein 73 | ||||
| TrEMBL | I1NQJ3 | 0.0 | I1NQJ3_ORYGL; Uncharacterized protein | ||||
| STRING | ORGLA01G0213300.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3075 | 37 | 86 | Representative plant | OGRP525 | 15 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G28500.1 | 1e-125 | NAC domain containing protein 73 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g48130.2 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



