 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g48060.1 | ||||||||
| Common Name | ARF2, LOC4325895, Os01g0670800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 723aa MW: 79112.9 Da PI: 7.1733 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.3 | 3e-23 | 147 | 248 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+lt sd++++g +++p++ ae++ +++ + ++l+ +d++g +W++++iyr++++r++lt+GW++F + ++L +gD v+F +
LOC_Os01g48060.1 147 FCKTLTASDTSTHGGFSVPRRAAEDCfppldySLQRPF-QELVAKDLHGTEWRFRHIYRGQPRRHLLTTGWSGFINKKKLVSGDAVLFL--RG 236
99****************************99777666.49************************************************..44 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
+++el+++v+r+
LOC_Os01g48060.1 237 EDGELRLGVRRA 248
9999*****996 PP
| |||||||
| 2 | Auxin_resp | 98.3 | 1.1e-32 | 273 | 357 | 1 | 82 |
Auxin_resp 1 aahaastksvFevvYnP....rastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsL 82
+aha++ ks+F+++YnP r s+seF+++++k+++++++++svGmRfk+++e+ed+serr +G+++g +++dp W++SkW++L
LOC_Os01g48060.1 273 VAHAVAVKSIFHIYYNPscthRLSQSEFIIPYWKFMRSFSQPFSVGMRFKLRYESEDASERRRTGIIIGSREADP-MWHGSKWKCL 357
799**************66666899**************************************************.9********9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.0E-38 | 139 | 252 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.44E-42 | 144 | 284 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.95E-21 | 145 | 247 | No hit | No description |
| PROSITE profile | PS50863 | 12.901 | 147 | 249 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.5E-21 | 147 | 248 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.5E-22 | 147 | 249 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.3E-26 | 273 | 357 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010050 | Biological Process | vegetative phase change | ||||
| GO:0010158 | Biological Process | abaxial cell fate specification | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0007042 | developmental stage | whole plant fruit formation stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 723 aa Download sequence Send to blast |
MVGIDLNTVE EEEDEEEGGA TGTVTAPAEA RAGGAVCLEL WHACAGPVAP LPRKGSAVVY 60 LPQGHLEHLG AAPGSGPGAA VPPHVFCRVV DVSLHADAAT DEVYAQVSLV ADNEEVERRM 120 REGEDGAACD GEGEDAVKRP ARIPHMFCKT LTASDTSTHG GFSVPRRAAE DCFPPLDYSL 180 QRPFQELVAK DLHGTEWRFR HIYRGQPRRH LLTTGWSGFI NKKKLVSGDA VLFLRGEDGE 240 LRLGVRRAAQ LKNASPFPAL HNQISNTSSL SEVAHAVAVK SIFHIYYNPS CTHRLSQSEF 300 IIPYWKFMRS FSQPFSVGMR FKLRYESEDA SERRRTGIII GSREADPMWH GSKWKCLVVK 360 WDDDVECRRP NGVSPWEIEL SGSVSGSHLS TPHSKRLKSC FPQVNPDIVL PNGSVSSDFA 420 ESARFHKVLQ GQELLGLKTR DGTVNTASQA TEARNFQYTD ERSCSINMSN NILGVPRLGV 480 KTPSGNPGFS YHCSGFGESQ RFQEVLQGQE VFRPYRGGTL SDACIRGSGF RQPDGNHAPG 540 AAFKWLAPQG CDHHGITTSV LPQASSPSSV LMFPQTSSKM PGLEYIYGCL DRNENSRHFK 600 IGPTQDMTRT DQTLRLWPHL ISGKVLDECT RNEKLHSPVS GAEHESNNKC LNTNGCKIFG 660 ISLTEKAQAG DEVDCGNASY HSRLQSLKPQ MPKSLGSSCA TVHEQRPVVG RVVDISAVNT 720 MI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldy_A | 1e-111 | 35 | 387 | 18 | 361 | Auxin response factor 1 |
| 4ldy_B | 1e-111 | 35 | 387 | 18 | 361 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.6367 | 0.0 | callus| flower| leaf| panicle| seed| stem| vegetative meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 19352034 | 0.0 | ||||
| Expression Atlas | Q0JKI9 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00021 | PBM | Transfer from AT2G33860 | Download |
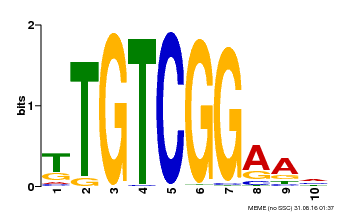
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g48060.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin under dark condition. {ECO:0000269|PubMed:17408882}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g48060 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB071291 | 0.0 | AB071291.1 Oryza sativa OsETTIN2 mRNA for Arabidopsis ETTIN-like protein 2, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015647434.1 | 0.0 | auxin response factor 2 isoform X1 | ||||
| Swissprot | Q0JKI9 | 0.0 | ARFB_ORYSJ; Auxin response factor 2 | ||||
| TrEMBL | I1NQJ0 | 0.0 | I1NQJ0_ORYGL; Auxin response factor | ||||
| STRING | ORGLA01G0213000.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2537 | 35 | 80 | Representative plant | OGRP6685 | 10 | 18 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33860.1 | 1e-162 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g48060.1 |
| Entrez Gene | 4325895 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



