 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os01g09640.1 | ||||||||
| Common Name | LOC4327305, Os01g0192300, OsJ_00711, OSNPB_010192300, P0671B11.19-1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 311aa MW: 33037.4 Da PI: 9.9691 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.6 | 3.8e-15 | 125 | 169 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++lG+g+W+ I+r++ ++Rt+ q+ s+ qky
LOC_Os01g09640.1 125 PWTEEEHRKFLVGLEKLGKGDWRGISRHFVTTRTPTQVASHAQKY 169
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50158 | 8.68 | 3 | 18 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 19.454 | 118 | 174 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.72E-19 | 119 | 175 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.4E-18 | 121 | 173 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 7.0E-12 | 122 | 172 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-11 | 122 | 168 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.26E-10 | 125 | 170 | No hit | No description |
| Pfam | PF00249 | 1.2E-12 | 125 | 169 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006338 | Biological Process | chromatin remodeling | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0035066 | Biological Process | positive regulation of histone acetylation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0003713 | Molecular Function | transcription coactivator activity | ||||
| GO:0004402 | Molecular Function | histone acetyltransferase activity | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000032 | anatomy | tetrad of microspores | ||||
| PO:0001029 | developmental stage | E tetrad stage | ||||
| PO:0007130 | developmental stage | sporophyte reproductive stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 311 aa Download sequence Send to blast |
MARKCSSCGN NGHNSRTCTG QRSLQESGGG YGGGGAGGVR LFGVQLHVGG APLKKCFSME 60 CLSSPSPSPS PAYYAAVAAA ASNSSPTVSS SSSLVSVEEA GEKMANGYLS DGLMARAQER 120 KKGVPWTEEE HRKFLVGLEK LGKGDWRGIS RHFVTTRTPT QVASHAQKYF LRQSSLTQKK 180 RRSSLFDVIE DAEKAPSVNE RLKLRHETAS VPAEMGFPAL SLGISSMAQP EAMLLPPPSL 240 TLTPSCSSPA VSSSSSEQPR TIHPSLMVAK PQVQLQLQPP DLELKISTVR QNDQPSSSPR 300 TPFLGTIRVT * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.623 | 0.0 | callus| flower| leaf| panicle| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32987612 | 0.0 | ||||
| Expression Atlas | Q5SNG6 | - | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
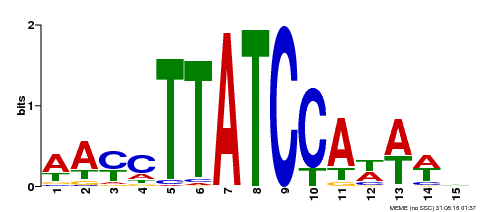
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os01g09640.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os01g09640 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK062109 | 0.0 | AK062109.1 Oryza sativa Japonica Group cDNA clone:001-045-B07, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015625837.1 | 0.0 | transcription factor MYBS3-like | ||||
| TrEMBL | Q5SNG6 | 0.0 | Q5SNG6_ORYSJ; Os01g0192300 protein | ||||
| STRING | OS01T0192300-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4231 | 38 | 73 | Representative plant | OGRP394 | 16 | 99 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56840.1 | 2e-49 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os01g09640.1 |
| Entrez Gene | 4327305 |



