 |
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
BGIOSGA036957-PA |
| Common Name | OsI_37401 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
| Family |
bZIP |
| Protein Properties |
Length: 783aa MW: 86089.8 Da PI: 5.3645 |
| Description |
bZIP family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| BGIOSGA036957-PA | genome | RIS | View CDS |
|
| Signature Domain? help Back to Top |
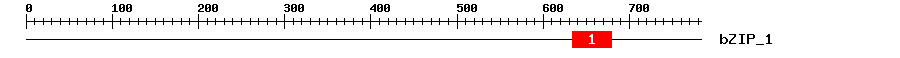 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | bZIP_1 | 30.4 | 8.6e-10 | 634 | 679 | 5 | 50 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkele 50
k+ r+ NReA r++Rq+Kka +++Lee+vk L N++L k+l+
BGIOSGA036957-PA 634 KKSRKPLGNREAVRKYRQKKKAHTAHLEEEVKRLRVINQQLVKRLQ 679
8999999***********************************9987 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated |
| GO:0000166 | Molecular Function | nucleotide binding |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0043565 | Molecular Function | sequence-specific DNA binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 783 aa
Download sequence Send
to blast |
MPRRTDNAAS ANSVEPEKSE ECLEFDDDDE EEVEEEEIEY EEIEEEIEEE EVEEDEDVVE 60
EVEEVDEEED EEEEEESDET EGVSKTKGVH QKDVTEKGKH AELLALPPHG SEVYVGGISS 120
DVSSEDLKRL CEPVGEVVEV RMMRGKDDSR GYAFVNFRTK GLALKAVKEL NNAKLKGKRI 180
RVSSSQAKNK LFIGNVPHSW TDDDFRKVVE EVGPGVLKAD LMKVSSANRN RGYGFVEYYN 240
HACAEYARQE MSSPTFKLDS NAPTVSWADP KNNDSASTSQ VKSVYVKNLP KNVTQAQLKR 300
LFEHHGEIEK VVLPPSRGGH DNRYGFVHFK DRSMAMRALQ NTERYELDGQ VLDCSLAKPP 360
AADKKDDRVP LPSSNGAPLL PSYPPLGYGI MSVPGAYGAA PASTAQPMLY APRAPPGAAM 420
VPMMLPDGRL VYVVQQPGGQ LPLASPPPQQ AGHRSGSGGR HGGSGGRYGG GGGSSGSSRP 480
EERVSETRLY MRILPVSQSA ESDGALYRQF TPVSHMDYEI SGPESEVDSE VWLGWVAQRR 540
RRRATAPYSG RRRVHEASDA GDDTAIRRSW QQVKEIPTSL DDFLPSIRTT CTHTHTCNPP 600
GPSATEHTHT CYHTHTRVFS SDDDSCGGDK AKPKKSRKPL GNREAVRKYR QKKKAHTAHL 660
EEEVKRLRVI NQQLVKRLQG QAALEVEVVR LRSLLVDVRS RINGALGSCP IQAQCGVDNV 720
LGCDGMAQCF AGKPELGVRQ SCAPSTVNCH ISSDSGQNLV VPHALSPSDV IGSFMVSSTS 780
KDE
|
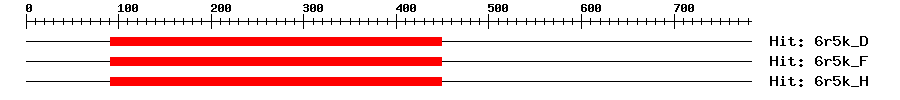
| 3D Structure ? help Back to Top |
 |
| PDB ID |
Evalue |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
| 6r5k_D | 4e-14 | 91 | 449 | 22 | 479 | Polyadenylate-binding protein, cytoplasmic and nuclear |
| 6r5k_F | 4e-14 | 91 | 449 | 22 | 479 | Polyadenylate-binding protein, cytoplasmic and nuclear |
| 6r5k_H | 4e-14 | 91 | 449 | 22 | 479 | Polyadenylate-binding protein, cytoplasmic and nuclear |
| Search in ModeBase |
| Nucleic Localization
Signal ? help
Back to Top |
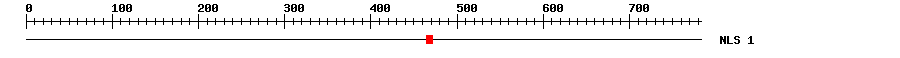 |
| No. |
Start |
End |
Sequence |
| 1 | 464 | 472 | GGRYGGGGG |
| Expression --
Description ? help
Back to Top |
| Source |
Description |
| Uniprot | DEVELOPMENTAL STAGE: In young plants, expressed in the distal regions of cotyledons, throughout leaves and root apical meristem, lateral root meristems and young floral buds. Strongly detected in the vascular tissues of various organs (e.g. roots, leaves, hypocotyls, sepals, petals, anther filaments) as well as in the gynophore. In the gynoecium, mainly detected in apical and basal regions as well as in the developing ovules in these regions. {ECO:0000269|PubMed:21304947}. |
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in vascular and meristematic tissues. Expressed throughout development in seedlings, roots, leaves, floral buds and siliques. {ECO:0000269|PubMed:21304947}. |
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Transcriptional activator that binds DNA on GAGA-like motif and 5'-(C/G)ACGTG(G/T)C(A/G)-3' consensus motif in the promoters of target genes (PubMed:27495811). Component of ribonucleosomes, which are complexes of at least 20 other different heterogenious nuclear ribonucleoproteins (hnRNP). hnRNP play an important role in processing of precursor mRNA in the nucleus (By similarity). Required during flower development and for cell fate determination (PubMed:21304947). Acts both as an antagonist and as a promoter of polycomb LHP1 gene regulation activity, depending of target genes, to regulate the transcription of stress-responsive and flowering genes (PubMed:21304947, PubMed:27495811). May regulate histone H3 trimethylation on lysine 27 (H3K27me3) (PubMed:21304947). Recognizes and binds histone H3 tails methylated at 'Lys-4' (H3K4me) and acetylated at 'Lys-9' (H3K9ac), leading to epigenetic activation. When in complex with LHP1, recognizes and binds histone H3 tails methylated at 'Lys-4' (H3K4me) and 'Lys-27' (H3K27me), mostly corresponding to stress-responsive genes (PubMed:27495811). May function as a suppressor of cell-autonomous immune responses involving glucosinolates, salicylic acid (SA) and jasmonic acid (JA) pathways toward pathogenic bacteria and fungi (PubMed:24914891). {ECO:0000250|UniProtKB:O60506, ECO:0000269|PubMed:21304947, ECO:0000269|PubMed:24914891, ECO:0000269|PubMed:27495811}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Slighty induced upon pathogen infection (e.g. P.syringae) (PubMed:24914891). Rapidly recruited to chromatin upon methyl jasmonate treatment (MeJA) to mediate transcriptional gene activation (PubMed:27495811). {ECO:0000269|PubMed:24914891, ECO:0000269|PubMed:27495811}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AK068700 | 0.0 | AK068700.1 Oryza sativa Japonica Group cDNA clone:J013160I09, full insert sequence. |
| GenBank | AK069493 | 0.0 | AK069493.1 Oryza sativa Japonica Group cDNA clone:J023024M07, full insert sequence. |




