 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA031911-PA | ||||||||
| Common Name | OsI_33602 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 476aa MW: 50786.9 Da PI: 7.8162 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.6 | 2.3e-05 | 106 | 128 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ C+k F r nL+ H r H
BGIOSGA031911-PA 106 FVCEVCNKGFQRDQNLQLHRRGH 128
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13 | 0.00031 | 191 | 213 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cgk++ +s+ k H +++
BGIOSGA031911-PA 191 WRCERCGKRYAVHSDWKAHVKNC 213
58******************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-5 | 105 | 128 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.28E-7 | 105 | 128 | No hit | No description |
| Pfam | PF12874 | 0.0017 | 106 | 128 | No hit | No description |
| PROSITE profile | PS50157 | 10.97 | 106 | 128 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.021 | 106 | 128 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 108 | 128 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 89 | 156 | 186 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.0E-4 | 179 | 211 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.28E-7 | 186 | 211 | No hit | No description |
| SMART | SM00355 | 99 | 191 | 211 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 476 aa Download sequence Send to blast |
MLLSDLSSDQ EATGSNSHGG GGGGDRMVVG SHGAAHVVLS NLFLPPAAAA AATMLLPAAP 60 VMVRPAAMAA AQEPRAKKKR SLPGNPDPEA EVIALSPRAL VATNRFVCEV CNKGFQRDQN 120 LQLHRRGHNL PWKLRHRAAA VSAVTTAAPA PRKRVYVCPE PTCVHHDPAR ALGDLTGIKK 180 HFSRKHGEKR WRCERCGKRY AVHSDWKAHV KNCGTREYRC DCGILFSRKD SLLTHRAFCD 240 ALAEESARLL AAANNSSSIT TTTCNNSNIS SNNNNNNINS ISNSNNLLIT SSSSSPPLFL 300 PFSTTPAENP NPNQLLFLQQ HQAAHHQLLL PQFQQPPSSP PAYFDHLAFG GGGGVITSSS 360 CNDDNSSIAG DVMVAAGGDS VSFGLTSEGS VTMHAGDVGR RRLTRDFLGV DHDAGEVDEL 420 ELDELPADLS TTAAACQGCN FAAATTVACC ATDFTTGSRQ YLGRLPPVNE TWSHNF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 1e-32 | 187 | 256 | 3 | 72 | Zinc finger protein JACKDAW |
| 5b3h_F | 1e-32 | 187 | 256 | 3 | 72 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.63282 | 0.0 | flower| panicle | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 200096392 | 0.0 | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that acts as a flowering master switch in both long and short days, independently of the circadian clock (By similarity). Promotes flowering upstream of HD1 by up-regulating FTL1, FTL4, FTL5, FTL6, EHD1, HD3A and RFT1 (By similarity). Seems to repress FTL11 expression (By similarity). May recognize the consensus motif 5'-TTTGTCGTAAT-3' in target gene promoters (By similarity). {ECO:0000250|UniProtKB:B1B534}. | |||||
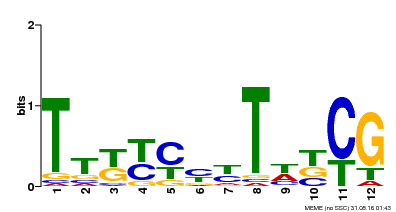
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00673 | SELEX | Transfer from GRMZM2G011357 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB359196 | 0.0 | AB359196.1 Oryza sativa Japonica Group Ehd2 mRNA for early heading date 2, complete cds, cultivar: Tohoku IL9. | |||
| GenBank | FJ009578 | 0.0 | FJ009578.1 Oryza sativa Japonica Group cultivar Zhonghua 11 Cys2/His2-type zinc finger transcription factor (RID1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015614407.1 | 0.0 | protein indeterminate-domain 12 | ||||
| Swissprot | B8BGV5 | 0.0 | EHD2_ORYSI; Protein EARLY HEADING DATE 2 | ||||
| TrEMBL | I1QUJ7 | 0.0 | I1QUJ7_ORYGL; Uncharacterized protein | ||||
| STRING | ORGLA10G0088000.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP13096 | 28 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G03840.1 | 4e-90 | C2H2 family protein | ||||



