 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA020087-PA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 329aa MW: 34943.2 Da PI: 6.9849 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 46.9 | 5.9e-15 | 259 | 302 | 5 | 48 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke 48
+r++r++kNRe+A rsR+RK+a++ eLe+kv Le+eN++Lkk+
BGIOSGA020087-PA 259 RRQKRMIKNRESAARSRARKQAYTNELENKVLRLEEENERLKKQ 302
79***************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.4E-14 | 253 | 302 | No hit | No description |
| SMART | SM00338 | 6.9E-10 | 255 | 311 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.553 | 257 | 302 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 8.82E-25 | 259 | 313 | No hit | No description |
| Pfam | PF00170 | 1.4E-12 | 259 | 302 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.13E-10 | 259 | 304 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 262 | 277 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 329 aa Download sequence Send to blast |
MASHAGGSGG GGGGSGRDAG SAQRGSMQGL ARQGSLYGLT LNEVQSQLGE PLLSMNLDEL 60 LKSVFPDGVD LDGGGGGIAG QSQPALGLQR QGSITMPPEL SKKTVDEVWK GIQDVPKRGA 120 EEGGRRRRER QPTLGEMTLE DFLVKAGVVT DPNDLPGNMD VVGGAAAAAA GTSDLNAGAQ 180 WLQQYHQQAL EPQHPSIGAP YMATHLAPQP LAVATGAVLD PIYSDGQITS PMLGALSDPQ 240 TPGRKRGATG EIADKLVERR QKRMIKNRES AARSRARKQA YTNELENKVL RLEEENERLK 300 KQKELDEILN AAPPPEPKYQ LRRTSSAAF |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 123 | 130 | GRRRRERQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.48133 | 0.0 | callus| panicle | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32973416 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in embryo during the latest stages of seed maturation. {ECO:0000269|PubMed:15642716}. | |||||
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in seeds. {ECO:0000269|PubMed:12376636}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the embryo specification element and the ABA-responsive element (ABRE) of the Dc3 gene promoter. Could participate in abscisic acid-regulated gene expression during seed development. | |||||
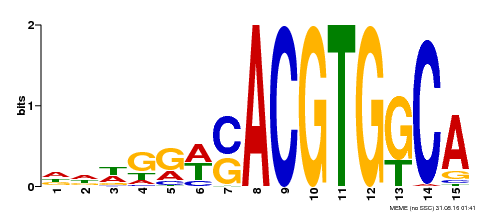
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00409 | DAP | Transfer from AT3G56850 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK063398 | 0.0 | AK063398.1 Oryza sativa Japonica Group cDNA clone:001-114-H11, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015638733.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 2 isoform X1 | ||||
| Swissprot | Q9LES3 | 1e-85 | AI5L2_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 2 | ||||
| TrEMBL | A0A0E0HG57 | 0.0 | A0A0E0HG57_ORYNI; Uncharacterized protein | ||||
| STRING | ONIVA05G21710.2 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2707 | 38 | 83 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G56850.1 | 3e-64 | ABA-responsive element binding protein 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



