 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA014158-PA | ||||||||
| Common Name | ARF11, ARF5, MP, OsI_017171, OSIGBa0099L20.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 926aa MW: 101965 Da PI: 6.1892 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 66.4 | 4e-21 | 143 | 230 | 1 | 83 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFk 83
f+k lt sd++++g +++p++ ae+ ++++++ ++l+++d++ + W++++iyr++++r++lt+GW+ Fv a++Lk+gD+v+F
BGIOSGA014158-PA 143 FCKNLTASDTSTHGGFSVPRRAAEKLfpqldySMQPPN-QELIVRDLHDNMWTFRHIYRGQPKRHLLTTGWSLFVGAKRLKAGDSVLFI 230
89*********************999*****9555555.49**********************************************96 PP
| |||||||
| 2 | Auxin_resp | 117.5 | 1.1e-38 | 240 | 322 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++s F+++YnPr+s+s Fv++v++++ka++++ svGmRf m+fete+ss+rr++Gtvvg+sd+dp+rWpnSkWr+L+
BGIOSGA014158-PA 240 AAHAASSGSSFTIYYNPRTSPSPFVIPVARYNKATYMQPSVGMRFAMMFETEESSKRRYTGTVVGISDYDPMRWPNSKWRNLQ 322
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 6.41E-39 | 133 | 236 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 8.1E-34 | 136 | 243 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.17E-19 | 142 | 229 | No hit | No description |
| SMART | SM01019 | 2.2E-16 | 143 | 248 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 7.8E-19 | 143 | 230 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.011 | 143 | 245 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.6E-34 | 240 | 322 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.9E-10 | 777 | 910 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 7.19E-9 | 820 | 902 | No hit | No description |
| PROSITE profile | PS51745 | 25.501 | 823 | 907 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 926 aa Download sequence Send to blast |
MASSQEKAKT GVLRNAAALL DEMQLMGETQ GAKKVINSEL WHACAGPLVC LPQRGSLVYY 60 FPQGHSEQVA ATTRKIPNSR IPNYPNLPSQ LLCQVHNITL HADKDTDEVY AQMTLQPVNS 120 ETDVFPIPTL GAYTKSKHPT EYFCKNLTAS DTSTHGGFSV PRRAAEKLFP QLDYSMQPPN 180 QELIVRDLHD NMWTFRHIYR GQPKRHLLTT GWSLFVGAKR LKAGDSVLFI SMHIGVLAAA 240 AHAASSGSSF TIYYNPRTSP SPFVIPVARY NKATYMQPSV GMRFAMMFET EESSKRRYTG 300 TVVGISDYDP MRWPNSKWRN LQVEWDEHGY GERPERVSIW DIETPENTLV FPSSTLNSKR 360 QCLPGYGVSV PGMEIGSANM SSFPRAQGNP YGSLQHIPAV GSELAIMLLN QSGQTLGSPL 420 SFHQSSYSSI IQNVKQNYIP PLTVSTSACL TKQESLPSDD AQHQFHMANM QNGDLEGSEV 480 QPVIDSISES KLNATSRDPR NTDSYTSRST SEQNSKGEPR GKTRRSKKGL PHKTVSEKSD 540 LSSAPSWICD NQQVGLESKL VGCDEQVNCG NIEDSSGALT QGNFVGQPHG HQVEQKGVLS 600 PPKVESSKSP DGGKSVNSFP NQGCFSQFID GLDWMTQPSY YQDSNVIQPA GVSENIFSSS 660 ADIPPSMIAD TMETFQASCL SDCLPNSIQE FISSPDLNSL TFLSPDMQNL EVQLQHDGSN 720 LPSTSNSFVQ MSFSEESASQ SANLSGLHME STHRSINTTS CSQPMSTGGF DAGMYSKLPR 780 LKESQILSLP EIHTNSMGTS ACSMDATEYS LDRSAKPMKP PVRTYTKVQK QGSVGRSIDV 840 TGFRNYHELR SAIACMFGLQ GKLEHPGSSE WKLVYVDYEN DVLLVGDDPW EEFINCVRCI 900 RILSPSEVQQ MSENGMHVLN DCIQAA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 1e-175 | 1 | 345 | 5 | 390 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.48513 | 0.0 | callus| panicle | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32988661 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
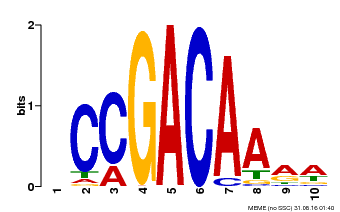
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB071292 | 0.0 | AB071292.1 Oryza sativa OsMP mRNA for Arabidopsis Monopteros-like protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025881052.1 | 0.0 | auxin response factor 11-like isoform X1 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A3AYC7 | 0.0 | A3AYC7_ORYSJ; Auxin response factor | ||||
| STRING | ORGLA04G0248600.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 1e-179 | ARF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



