 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA005099-PA | ||||||||
| Common Name | OsI_05027 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 803aa MW: 90387.2 Da PI: 6.4775 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 69.6 | 4.1e-22 | 124 | 225 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldg 86
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d+++ W++++i+r++++r++l++GW+ Fv++++L +gD ++F +
BGIOSGA005099-PA 124 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHSMDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--R 212
99*****************************7555544..58************************************************..4 PP
SSEE..EEEEE-S CS
B3 87 rsefelvvkvfrk 99
+++el+v+v+r+
BGIOSGA005099-PA 213 GENGELRVGVRRA 225
49999*****996 PP
| |||||||
| 2 | Auxin_resp | 113.6 | 1.9e-37 | 250 | 331 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++svGmRf+m+fe+e+++e+r++Gt++g ++ldp Wp+S WrsLk
BGIOSGA005099-PA 250 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFEGEEAPEQRFTGTIIGSENLDP-VWPESSWRSLK 331
89*********************************************************************.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 8.5E-40 | 116 | 253 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.3E-39 | 117 | 239 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.30E-19 | 122 | 224 | No hit | No description |
| SMART | SM01019 | 1.5E-21 | 124 | 226 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.265 | 124 | 226 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.0E-20 | 124 | 225 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.8E-34 | 250 | 331 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 8.4E-7 | 668 | 723 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 4.46E-13 | 683 | 764 | No hit | No description |
| PROSITE profile | PS51745 | 26.238 | 687 | 780 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 9.7E-4 | 705 | 760 | No hit | No description |
| Pfam | PF02309 | 2.6E-8 | 729 | 772 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 803 aa Download sequence Send to blast |
MAPPPPPQGS STGDPLYDEL WHACAGPLVT VPRVGDLVFY FPQGHIEQVE ASMNQVADSQ 60 MRLYDLPSKL LCRVLNVELK AEQDTDEVYA QVMLMPEPEQ NEMAVEKTTP TSGPVQARPP 120 VRSFCKTLTA SDTSTHGGFS VLRRHADECL PPLDMTQSPP TQELVAKDLH SMDWRFRHIF 180 RGQPRRHLLQ SGWSVFVSSK RLVAGDAFIF LRGENGELRV GVRRAMRQLS NVPSSVISSQ 240 SMHLGVLATA WHAINTKSMF TVYYKPRTSP SEFIIPYDQY MESVKNNYSV GMRFRMRFEG 300 EEAPEQRFTG TIIGSENLDP VWPESSWRSL KVRWDEPSTI PRPDRVSPWK IEPASSPPVN 360 PLPLSRVKRP RPNAPPASPE SPILTKEAAT KVDTDPAQAQ RSQNSTVLQG QEQMTLRSNL 420 TESNDSDVTA HKPMMWSPSP NAAKAHPLTF QQRPPMDNWM QLGRRETDFK DVRSGSQSFG 480 DSPGFFMQNF DEAPNRLTSF KNQFQDQGSA RHFSDPYYYV SPQPSLTVES STQMHTDSKE 540 LHFWNGQSTV YGNSRDRPQN FRFEQNSSSW LNQSFARPEQ PRVIRPHASI APVELEKTEG 600 SGFKIFGFKV DTTNAPNNHL SSPMAATHEP MLQTPSSLNQ LQPVQTDCIP EVSVSTAGTA 660 TENEKSGQQA QQSSKDVQSK TQVASTRSCT KVHKQGVALG RSVDLSKFSN YDELKAELDK 720 MFEFDGELVS SNKNWQIVYT DNEGDMMLVG DDPWEEFCSI VRKIYIYTKE EVQKMNSKSN 780 APRKDDSSEN EKGHLPMPNK SDN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-169 | 9 | 356 | 12 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-169 | 9 | 356 | 12 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-169 | 9 | 356 | 12 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-169 | 9 | 356 | 12 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-169 | 9 | 356 | 12 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 367 | 371 | KRPRP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.26144 | 0.0 | callus| flower| leaf| panicle| root| seed| stem| vegetative meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32982332 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
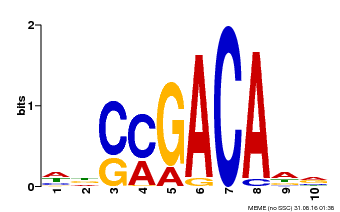
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072309 | 0.0 | AK072309.1 Oryza sativa Japonica Group cDNA clone:J023022C23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015650953.1 | 0.0 | auxin response factor 4 | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | B8A8I4 | 0.0 | B8A8I4_ORYSI; Auxin response factor | ||||
| STRING | OS01T0927600-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



