 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA004670-PA | ||||||||
| Common Name | GAM1, OsI_004080 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 553aa MW: 59960.7 Da PI: 4.8859 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.5 | 1.1e-16 | 42 | 89 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+vk++G g+W+++ + g+ R++k+c++rw ++l
BGIOSGA004670-PA 42 KGPWTSAEDAILVDYVKKHGEGNWNAVQKNTGLFRCGKSCRLRWANHL 89
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 51 | 3.4e-16 | 95 | 138 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l+++++ ++G++ W++ a++++ gRt++++k++w++
BGIOSGA004670-PA 95 KGAFTAEEERLIIQLHSKMGNK-WARMAAHLP-GRTDNEIKNYWNT 138
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF001693 | 1.3E-297 | 1 | 553 | IPR016310 | Transcription factor, GAMYB |
| PROSITE profile | PS51294 | 16.625 | 37 | 89 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.49E-31 | 39 | 136 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-14 | 41 | 91 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-14 | 42 | 89 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-23 | 43 | 96 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.69E-11 | 44 | 89 | No hit | No description |
| PROSITE profile | PS51294 | 25.026 | 90 | 144 | IPR017930 | Myb domain |
| SMART | SM00717 | 6.3E-17 | 94 | 142 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.8E-15 | 95 | 138 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.20E-12 | 97 | 138 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-25 | 97 | 142 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 553 aa Download sequence Send to blast |
MYRVKSESDC EMIHQEQMDS PVADDGSSGG SPHRGGGPPL KKGPWTSAED AILVDYVKKH 60 GEGNWNAVQK NTGLFRCGKS CRLRWANHLR PNLKKGAFTA EEERLIIQLH SKMGNKWARM 120 AAHLPGRTDN EIKNYWNTRI KRCQRAGLPI YPTSVCNQSS NEDQQCSSDF DCGENLSNDL 180 LNANGLYLPD FTCDNFIANS EALPYAPHLS AVSISNLLGQ SFASKSCSFM DQVNQTGMLK 240 QSDGVLPGLS DTINGVISSV DQFSNDSEKL KQAVGFDYLH EANSTSKIIA PFGGALNGSH 300 AFLNGNFSAS RPTSGPLKME LPSLQDTESD PNSWLKYTVA PALQPTELVD PYLQSPAATP 360 SVKSECASPR NSGLLEELVH EAQTLRSGKN QQTSVISSSS SVGTPCNTTV LSPEFDMCQE 420 YWEEQHPGPF LNDCAPFSGN SFTESTPPVS AASPDIFQLS KVSPAQSTSM GSGEQVMGPK 480 YEPGDTSPHP ENFRPDALFS GNTADPSVFN NAIAMLLGND LSIDCRPVLG DGIMFNSSSW 540 SNMPHACEMS EFK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 7e-33 | 36 | 142 | 1 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.2698 | 0.0 | callus| flower| panicle| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32973969 | 0.0 | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
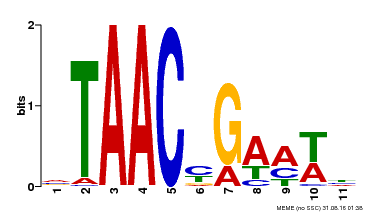
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CT830882 | 0.0 | CT830882.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCFA246D08, full insert sequence. | |||
| GenBank | CT830884 | 0.0 | CT830884.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCFA248K02, full insert sequence. | |||
| GenBank | X98355 | 0.0 | X98355.1 O.sativa mRNA for transcription activator, GAMyb. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015622335.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_015622336.1 | 0.0 | transcription factor GAMYB | ||||
| Swissprot | A2WW87 | 0.0 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A0D3EVB7 | 0.0 | A0A0D3EVB7_9ORYZ; Transcription factor | ||||
| STRING | OBART01G34660.2 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3947 | 36 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 6e-88 | myb domain protein 33 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



