 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA000374-PA | ||||||||
| Common Name | OsI_04690 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 303aa MW: 32934.7 Da PI: 8.8616 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 165.5 | 1.9e-51 | 9 | 133 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
lppGfrFhPtdeelv++yL+++++g ++ + +i+e+d+yk++Pw+Lp+++ +ekewyfFs+rd+ky++g+r+nra+ sgyWkatg dk+v
BGIOSGA000374-PA 9 LPPGFRFHPTDEELVMHYLCRRCAGLPIAV-PIIAEIDLYKFDPWQLPRMALYGEKEWYFFSPRDRKYPNGSRPNRAAGSGYWKATGADKPVG 100
79**************************99.88***************7666789*************************************9 PP
NAM 94 skkgelvglkktLvfykgrapkgektdWvmheyrl 128
s + v +kk Lvfy g+apkgekt+W+mheyrl
BGIOSGA000374-PA 101 S--PKPVAIKKALVFYAGKAPKGEKTNWIMHEYRL 133
9..789***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 7.85E-62 | 6 | 159 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 59.747 | 9 | 159 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.2E-26 | 10 | 133 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 303 aa Download sequence Send to blast |
MSGGQDLQLP PGFRFHPTDE ELVMHYLCRR CAGLPIAVPI IAEIDLYKFD PWQLPRMALY 60 GEKEWYFFSP RDRKYPNGSR PNRAAGSGYW KATGADKPVG SPKPVAIKKA LVFYAGKAPK 120 GEKTNWIMHE YRLADVDRSA RKKNSLRLDD WVLCRIYNKK GGLEKPPAAA VAAAGMVSNG 180 GGVERKPMVG VNAAVSSPPE QKPVVAGPAF PDLAAYYDRP SDSMPRLHAD SSCSEQVLSP 240 EFACEVQSQP KISEWERTFA TVGPINPAAS ILDPAGSGGL GGLGGGGSDP LLQDILMYWG 300 KPF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 1e-78 | 6 | 165 | 12 | 174 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Widely expressed. {ECO:0000269|PubMed:10660065}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the promoter of the stress response gene LEA19. Involved in tolerance to abiotic stresses (PubMed:20632034). Transcription activator involved in response to abiotic and biotic stresses. Involved in drought and salt stress responses, and defense response to the rice blast fungus (PubMed:17587305). Transcription activator involved tolerance to cold and salt stresses (PubMed:18273684). Transcription activator involved in tolerance to drought stress. Targets directly and activates genes involved in membrane modification, nicotianamine (NA) biosynthesis, glutathione relocation, accumulation of phosphoadenosine phosphosulfate and glycosylation in roots (PubMed:27892643). Controls root growth at early vegetative stage through chromatin modification and histone lysine deacytaltion by HDAC1 (PubMed:19453457). {ECO:0000269|PubMed:17587305, ECO:0000269|PubMed:18273684, ECO:0000269|PubMed:19453457, ECO:0000269|PubMed:20632034, ECO:0000269|PubMed:27892643}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00121 | DAP | Transfer from AT1G01720 | Download |
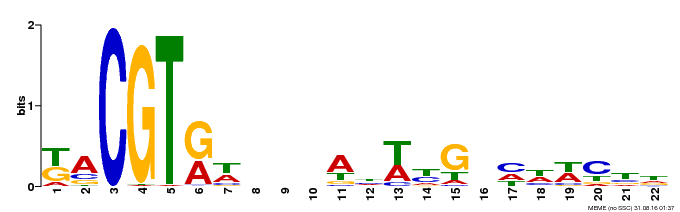
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by drought stress, salt stress, cold stress and abscisic acid (ABA) (PubMed:20632034, PubMed:27892643). Induced by methyl jasmonate (PubMed:20632034, PubMed:11332734). Induced by infection with the rice blast fungus Magnaporthe oryzae (PubMed:11332734). {ECO:0000269|PubMed:11332734, ECO:0000269|PubMed:20632034, ECO:0000269|PubMed:27892643}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB028185 | 0.0 | AB028185.1 Oryza sativa mRNA for OsNAC6 protein, complete cds. | |||
| GenBank | AK068392 | 0.0 | AK068392.1 Oryza sativa Japonica Group cDNA clone:J013149P14, full insert sequence. | |||
| GenBank | EU846993 | 0.0 | EU846993.1 Oryza sativa Japonica Group clone KCS089G05 NAC6 protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015620920.1 | 0.0 | NAC domain-containing protein 48 | ||||
| Swissprot | Q7F2L3 | 0.0 | NAC48_ORYSJ; NAC domain-containing protein 48 | ||||
| TrEMBL | A2WXN9 | 0.0 | A2WXN9_ORYSI; Uncharacterized protein | ||||
| STRING | ONIVA01G44460.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3417 | 37 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01720.1 | 1e-121 | NAC family protein | ||||



