 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI04G22600.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 611aa MW: 64301.2 Da PI: 6.0795 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 15.1 | 6.6e-05 | 165 | 182 | 2 | 19 |
EETTTTEEESSHHHHHHH CS
zf-C2H2 2 kCpdCgksFsrksnLkrH 19
+C++Cgk F++++++ H
ORUFI04G22600.1 165 VCKNCGKEFTSWEHFLEH 182
7**************999 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 9.536 | 76 | 98 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.56E-6 | 76 | 98 | No hit | No description |
| SMART | SM00355 | 0.96 | 76 | 98 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF13912 | 0.0051 | 77 | 98 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 78 | 98 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.56E-6 | 159 | 183 | No hit | No description |
| SMART | SM00355 | 140 | 164 | 184 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.141 | 164 | 191 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.58E-8 | 322 | 345 | No hit | No description |
| Pfam | PF13912 | 3.5E-9 | 323 | 347 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 10.242 | 323 | 350 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.13 | 323 | 345 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 325 | 345 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.58E-8 | 431 | 458 | No hit | No description |
| Pfam | PF13912 | 5.2E-11 | 436 | 459 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.7 | 436 | 458 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.182 | 436 | 458 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 438 | 458 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 611 aa Download sequence Send to blast |
MAVLDLQVQE NSRHCCDREF VTPDGGAHLI GAPISRSNSI PPRRTKRMHH HGQMQPSPSS 60 PRAPPPTAAQ ASGYKHFCRV CNKGFTCGSA LGGHMRAHGV GDGDGLGADD DDDDDDDSLG 120 DEAVRRARGG ADDPWNAGGP SSSGAATHVY ELRTNPNRVT RSRQVCKNCG KEFTSWEHFL 180 EHGKCSSGED DDDEDDVDRS LQPWSPSPEA DGEEDPAPAA GWLKGKRSRR CKGTGVDLSP 240 TPSACAAGEE EDLANCLVML SSSKVDQAGV TEAEQRSSSS ASKEHKRLIT FMEPTTYVLD 300 TVMALPPPAP APQYVSTVPR GMFECKACKK VFSSHQALGG HRASHKKVKG CFAAKLESNA 360 AEVAEPSHHA EVADRSEDNP AKATSDARRN VHASIDGDGN AGTSDAAAEL SMAIVPIEPP 420 VAALAAAPLK KKGKMHECSV CHRLFTSGQA LGGHKRCHWL TSSSADHTAS VPPLADDLVP 480 LSFRPMLDAP EPALDLSIAA NPPLLASAAT VRPKVGGSSF HLDAPPPVYI PSSPAIPSQR 540 NKATATTGSQ NANDAVGLST AAAEDEADST TVKRARLSDL KDVSMAGETT PWLQVGIGSS 600 SRGGADDNDK E |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 428 | 434 | LKKKGKM |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00621 | PBM | Transfer from AT2G17180 | Download |
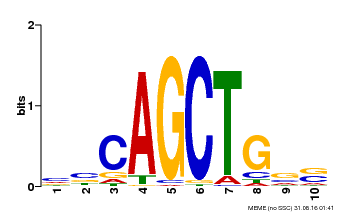
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP014960 | 0.0 | AP014960.1 Oryza sativa Japonica Group DNA, chromosome 4, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015634601.1 | 0.0 | uncharacterized protein LOC4336604 | ||||
| TrEMBL | A0A0E0PCE4 | 0.0 | A0A0E0PCE4_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI04G22600.1 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3524 | 32 | 62 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G17180.1 | 1e-20 | C2H2-like zinc finger protein | ||||



