 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI04G19390.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 619aa MW: 63352.1 Da PI: 6.8225 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 33.5 | 1.1e-10 | 281 | 337 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + ++ k + g ++ +++Aa+a++ a++k++g
ORUFI04G19390.1 281 SIYRGVTRHRWTGRYEAHLWDnSCR-RegQSrKGRQ-GGYDKEDKAARAYDLAALKYWG 337
57*******************5444.2455535555.7799999*************98 PP
| |||||||
| 2 | AP2 | 51.3 | 2.8e-16 | 380 | 431 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV+++++ grW A+I + +k ++lg+f t+eeAa+a++ a+ k++g
ORUFI04G19390.1 380 SKYRGVTRHHQHGRWQARIGRVAG---NKDIYLGTFSTEEEAAEAYDIAAIKFRG 431
89****************988532...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 3.2E-8 | 281 | 337 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.75E-14 | 281 | 347 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 4.42E-16 | 281 | 347 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 2.6E-11 | 282 | 347 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.7E-22 | 282 | 351 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 17.108 | 282 | 345 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.0E-6 | 283 | 294 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.22E-18 | 380 | 441 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 3.60E-23 | 380 | 441 | No hit | No description |
| Pfam | PF00847 | 1.6E-10 | 380 | 431 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.9E-18 | 381 | 440 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.401 | 381 | 439 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.3E-34 | 381 | 445 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.0E-6 | 421 | 441 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000723 | Biological Process | telomere maintenance | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0010073 | Biological Process | meristem maintenance | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0019827 | Biological Process | stem cell population maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 619 aa Download sequence Send to blast |
MASADNWLGF SLSGQGNPQH HQNGSPSAAG DAAIDISGSG DFYGLPTPDA HHIGMAGEDA 60 PYGVMDAFNR GTHETQDWAM RGLDYGGGSS DLSMLVGSSG GGRRTVAGDG GGEAPKLENF 120 LDGNSFSDVH GQAAGGYLYS GSAVGGAGGY SNGGCGGGTI ELSMIKTWLR SNQSQQQPSP 180 PQHADQGMST DASASSYACS DVLVGSCGGG GGGGAGGTAS SHGQGLALSM STGSVAAAAG 240 GGAVVAAESS SSENKRVDSP GGAVDGAVPR KSIDTFGQRT SIYRGVTRHR WTGRYEAHLW 300 DNSCRREGQS RKGRQGGYDK EDKAARAYDL AALKYWGTTT TTNFPMSNYE KELEEMKHMT 360 RQEYIAHLRR NSSGFSRGAS KYRGVTRHHQ HGRWQARIGR VAGNKDIYLG TFSTEEEAAE 420 AYDIAAIKFR GLNAVTNFDM SRYDVKSILD SSTLPVGGAA RRLKEAEAAA AAAGGGVIVS 480 HLADGGVGGY YYGCGPTIAF GGGGQQPAPL AVHYPSYGQA SGWCKPEQDA VIAAGHCATD 540 LQHLHLGSGG AAATHNFFQQ PASSSAVYGN GGGGGGNAFM MPMGAVVAAA DHGGQSSAYG 600 GGGEGSSWGT TASSTRTRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that promotes cell proliferation, differentiation and morphogenesis, especially during embryogenesis. {ECO:0000269|PubMed:12172019}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00368 | DAP | Transfer from AT3G20840 | Download |
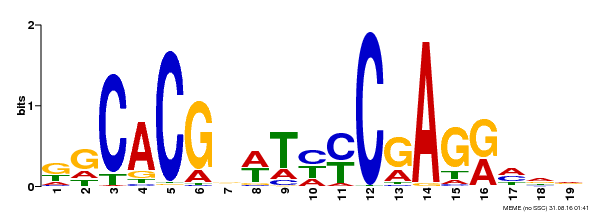
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK287726 | 0.0 | AK287726.1 Oryza sativa Japonica Group cDNA, clone: J065138E17, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015635553.1 | 0.0 | AP2-like ethylene-responsive transcription factor BBM1 | ||||
| Swissprot | Q8LSN2 | 1e-131 | BBM2_BRANA; AP2-like ethylene-responsive transcription factor BBM2 | ||||
| TrEMBL | A0A0E0PB96 | 0.0 | A0A0E0PB96_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI04G19390.1 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1569 | 37 | 110 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G20840.1 | 1e-115 | AP2 family protein | ||||



