 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI01G45560.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 412aa MW: 42654.3 Da PI: 9.0874 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.4 | 2.6e-40 | 92 | 169 | 2 | 78 |
-SSTT-----TT-.-HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlsea.keyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C vegC+adls++ ++yhrrhkvCe+hsk++vv+v+g++qrfCqqCsrfh l efDeekrsCr+rL++hn+rrrk+q+
ORUFI01G45560.1 92 CSVEGCAADLSKCvRDYHRRHKVCEAHSKTAVVTVAGQQQRFCQQCSRFHLLGEFDEEKRSCRKRLDGHNKRRRKPQP 169
***********9879************************************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.7E-33 | 84 | 154 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.487 | 89 | 167 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.88E-37 | 90 | 172 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.3E-29 | 92 | 166 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 412 aa Download sequence Send to blast |
MDWDAKMPSW DLGTVVGPSG GGGGGGGGGG ALDLKLGAPT SWKTTTTVSA ASAAPAAVAP 60 PPPPPASSSS SAAAAGKRAR AGQGQQAAVP ACSVEGCAAD LSKCVRDYHR RHKVCEAHSK 120 TAVVTVAGQQ QRFCQQCSRF HLLGEFDEEK RSCRKRLDGH NKRRRKPQPD PLNPGNLFAN 180 HHGAARFTSY PQIFSTAASM SPQETKWPAN VVKTEAADVF QEPYYHALHL NGAGAAAAAS 240 IFHHGGNKAR KHHFPFLTAD HGGGAAAASP LFGCQPFTIT PSSESRSSSS SRHSNGKMFA 300 HDGGLDNCAL SLLSDNPTPT AQITIPQPLF AGGGQYGGGG GGDVSLTGLS YVRMAGKDTS 360 ILAKSATTTA TTATTPTTTS AQLQYHGYYH HHVSADQGSS DAAIQALPFS SW |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-27 | 82 | 166 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 149 | 166 | KRSCRKRLDGHNKRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
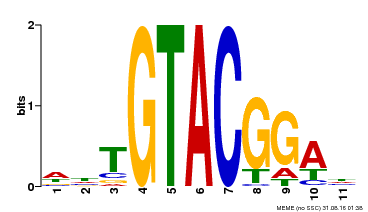
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK062581 | 0.0 | AK062581.1 Oryza sativa Japonica Group cDNA clone:001-100-B10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015611358.1 | 0.0 | squamosa promoter-binding-like protein 2 | ||||
| Swissprot | Q0JGI1 | 0.0 | SPL2_ORYSJ; Squamosa promoter-binding-like protein 2 | ||||
| TrEMBL | A0A0E0N6Y2 | 0.0 | A0A0E0N6Y2_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI01G45560.1 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10456 | 30 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 7e-30 | SBP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



