 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | e_gw1.7.793.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 335aa MW: 36033 Da PI: 4.6636 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 53.6 | 3.1e-17 | 300 | 331 | 1 | 32 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrk 32
C +Cgt kTp+WR gp+g+ktLCnaCG+++ k
e_gw1.7.793.1 300 CLHCGTVKTPQWRMGPEGKKTLCNACGVRFMK 331
99***************************977 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00401 | 6.5E-6 | 294 | 334 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 11.873 | 294 | 327 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 3.18E-12 | 295 | 331 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 9.8E-14 | 299 | 331 | IPR013088 | Zinc finger, NHR/GATA-type |
| Pfam | PF00320 | 9.9E-15 | 300 | 331 | IPR000679 | Zinc finger, GATA-type |
| CDD | cd00202 | 2.08E-12 | 300 | 329 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 300 | 325 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 335 aa Download sequence Send to blast |
MVTGERGTTT TTEDVGGASA AREAEDEPTP PATDEAWDVI NSFPIVRGDM VMCAGDGEQS 60 YGVNAPFIPP EFIERPIAAE AEGSDNEAEE DEEEDARRFS QAIRESSLLC EDDDVFDDDI 120 GAHADFISEP VAFAARREAR ELAAAAQVPG IDFMVKSGAL LCDDDYDAKD GNLECSAMTN 180 VSDYASEEEH FATAADGRVF RVSQALVAPS ASKLAKGHGK HRLEDSANRG GNDSEDSDAD 240 LVVQSARKAV PGKKTLATKR KRRPDAPIRR VKAAKKEKGS SVRATDIYVM SNGKRQRRGC 300 LHCGTVKTPQ WRMGPEGKKT LCNACGVRFM KGIL* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00530 | DAP | Transfer from AT5G25830 | Download |
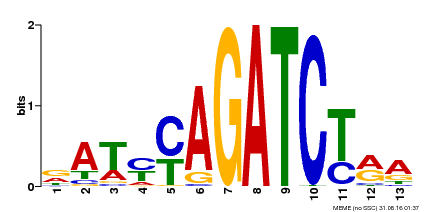
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001418973.1 | 6e-85 | predicted protein | ||||
| TrEMBL | A4S0H6 | 1e-83 | A4S0H6_OSTLU; Uncharacterized protein | ||||
| STRING | ABO97266 | 2e-84 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G25830.1 | 1e-15 | GATA transcription factor 12 | ||||



