 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | e_gw1.1.649.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 306aa MW: 33407.7 Da PI: 8.7424 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 93.1 | 2.3e-29 | 141 | 194 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+peLH++Fv av+qL G++kA+Pk+il+lm+v+gLt+e+v+SHLQkYRl
e_gw1.1.649.1 141 KPRVVWSPELHAQFVTAVNQL-GIDKAVPKRILDLMGVQGLTRENVASHLQKYRL 194
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 40.1 | 1.9e-14 | 1 | 56 | 53 | 109 |
CTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 53 mpgmdGlellkeireeepklpiivvtahgeeedalealkaGakdflsKpfdpeelvk 109
mp+mdG++ll+ i++e ++p+++++a+++++ +l+ + Ga d+l Kp+ +eel++
e_gw1.1.649.1 1 MPDMDGFKLLEIIQFEL-SIPVLMMSANSDSSVVLRGIIHGAVDYLLKPVRIEELRN 56
9************9977.*************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00156 | 2.47E-15 | 1 | 61 | No hit | No description |
| PROSITE profile | PS50110 | 24.823 | 1 | 62 | IPR001789 | Signal transduction response regulator, receiver domain |
| Gene3D | G3DSA:3.40.50.2300 | 7.2E-19 | 1 | 85 | No hit | No description |
| Pfam | PF00072 | 1.9E-10 | 1 | 57 | IPR001789 | Signal transduction response regulator, receiver domain |
| SuperFamily | SSF52172 | 3.98E-16 | 1 | 68 | IPR011006 | CheY-like superfamily |
| PROSITE profile | PS51294 | 12.203 | 138 | 197 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.7E-31 | 139 | 199 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.58E-20 | 139 | 198 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.7E-25 | 141 | 194 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.2E-8 | 143 | 193 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 306 aa Download sequence Send to blast |
MPDMDGFKLL EIIQFELSIP VLMMSANSDS SVVLRGIIHG AVDYLLKPVR IEELRNIWQH 60 IVRRDYTTKS PGSDEATTSS PNKRLKLSNS KSDEIDRSMN EAASSSKRKK PAKKGGKTGK 120 ESEKREEGAE KKSAEGAGAK KPRVVWSPEL HAQFVTAVNQ LGIDKAVPKR ILDLMGVQGL 180 TRENVASHLQ KYRLYLKRIQ GTESNSRNGT NAARHASNDG MPQSRPKDVP EKSTKTRAEA 240 FSADDTIFRG DDLSLPGDLL VGDEVGYDDP GFDPALLTPL DIDKVGKGDG LGATDQMLNF 300 FLSGD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5lxu_A | 6e-23 | 141 | 197 | 1 | 57 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
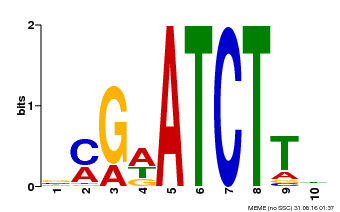
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022840333.1 | 1e-124 | Signal transduction response regulator, receiver domain | ||||
| Swissprot | B8AEH1 | 8e-65 | ORR23_ORYSI; Two-component response regulator ORR23 | ||||
| TrEMBL | A0A1Y5HZJ9 | 1e-130 | A0A1Y5HZJ9_OSTTA; Uncharacterized protein | ||||
| STRING | A0A096P7X7 | 1e-124 | (Ostreococcus tauri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP1220 | 15 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 3e-55 | response regulator 2 | ||||



