 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC08G03280.6 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 839aa MW: 92167.3 Da PI: 6.9793 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.6 | 3.9e-15 | 115 | 159 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
OPUNC08G03280.6 115 RERWTEAEHKRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 159
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.51E-16 | 109 | 165 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.147 | 110 | 164 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.0E-16 | 113 | 162 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.5E-12 | 114 | 162 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.5E-8 | 115 | 155 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.8E-12 | 115 | 158 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.28E-8 | 117 | 160 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MARPVTAAVA GCSGTRFGSG WLVDPLRFGW GATQAAVWRA AGVCGAERAS ELGWDCFACS 60 RGRRRLAFVR WWCCARFKEV FGLSRLTAIG SIQINSSGEE AVVKVRKPYT ITKQRERWTE 120 AEHKRFLEAL KLYGRAWQRI EEHVGTKTAV QIRSHAQKFF TKLEKEAINN GTSPGQAHDI 180 DIPPPRPKRK PNSPYPRKCS LSSETSTREV PNDKTTKSNM TNNSTAQMAG DAVLEKLQRK 240 EISEKGSCSE VLNLFREVPS ASFSSVNKSS SNHGASRGLE PTKIEVKDVV ILERDSISNG 300 AGKDVKDIDD QEMERLNGIH ISSKTDHSHE NCLDTSRQQF KPKSNSVETT YVDWSAVKAS 360 HYQMDRNGVT DFQATGTEGS HPDQTSDQMG ASGTMNQCIH PTLSVDPKFD GNATAQPFPH 420 NYAAFAPMIQ CHCNQDAYRS FANMSSTFSS MLVSTLLSNP AIHAAARLAA SYWPTVDSNT 480 PDPNQENLSE SAQGSHAGSP PNMASIVAAT VAAASAWWAT QGLLPLFPPP IAFPFVPAPT 540 APFSTADVQR AQEKDIDCPI DNAQKELQET RKQDKSEAMK VIVSSETDES GKGEVSLHTE 600 LKISPADKAD TKPATGADTS DVFGNKKKQD RSSCGSNTPS SSDIEADNAP EKQEKANDKA 660 NQASCSNSSA GDNNHRRFRS SASTSDSWKE VSEEVVVYQH FAYSHLSYSL NIHHCFSNSH 720 FCCQGRLAFD ALFSRERLPQ SFSPPQVEGS KEISKEEEDE VTTVTVDLNK NATIIDQELD 780 TVDEPRASFP NELSNLKLKS RRTGFKPYKR CSVEAKENRV PASDEVGTKR IRLESETST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
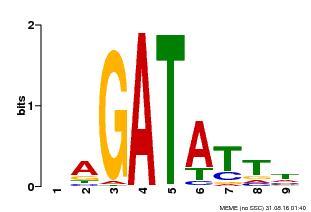
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004460 | 0.0 | AP004460.2 Oryza sativa Japonica Group genomic DNA, chromosome 8, PAC clone:P0438H08. | |||
| GenBank | AP014964 | 0.0 | AP014964.1 Oryza sativa Japonica Group DNA, chromosome 8, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015649844.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_015649845.1 | 0.0 | protein LHY isoform X2 | ||||
| TrEMBL | A0A0E0LRF7 | 0.0 | A0A0E0LRF7_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC08G03280.6 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.3 | 8e-39 | MYB_related family protein | ||||



