 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC05G22370.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 399aa MW: 41939.5 Da PI: 10.2063 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.5 | 1.4e-12 | 73 | 121 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ r +r++lg+f + eAa+a++ a+++++g
OPUNC05G22370.1 73 SKYKGVVPQP-NGRWGAQIYE------RhQRVWLGTFTGEAEAARAYDVAAQRFRG 121
89****9888.8*********......44**********99*************98 PP
| |||||||
| 2 | B3 | 91.9 | 4.6e-29 | 202 | 310 | 1 | 91 |
EEEE-..-HHHHTT-EE--HHH.HTT..................---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--T CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh..................ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLke 76
f+k+ tpsdv+k++rlv+pk++ae+h g + +++ l +ed g++W+++++y+++s++yvltkGW++Fvk++gL +
OPUNC05G22370.1 202 FDKTVTPSDVGKLNRLVIPKQHAEKHfplqipppttsvvadagaGAGECKGVLLNFEDAAGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKGLHA 295
89***************************88888888877777744455999****************************************** PP
T-EEEEEE-SSSEE. CS
B3 77 gDfvvFkldgrsefe 91
gD v F++ + ++
OPUNC05G22370.1 296 GDAVGFYRAAGKHAQ 310
*******76545554 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.7E-8 | 73 | 121 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.10E-22 | 73 | 129 | No hit | No description |
| SuperFamily | SSF54171 | 1.7E-16 | 73 | 129 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 3.2E-26 | 74 | 135 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.862 | 74 | 129 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.5E-19 | 74 | 129 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 6.3E-40 | 193 | 324 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.75E-27 | 200 | 303 | No hit | No description |
| SuperFamily | SSF101936 | 1.96E-28 | 200 | 316 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 2.4E-26 | 202 | 313 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.9E-20 | 202 | 318 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.153 | 202 | 319 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 399 aa Download sequence Send to blast |
MDSTSCLLDD ASSGASTGKK APAPAAPAAT TSKALQRVGS GASAVMDAAE PGAEADSGGE 60 RRAGGGGGKL PSSKYKGVVP QPNGRWGAQI YERHQRVWLG TFTGEAEAAR AYDVAAQRFR 120 GRDAVTNFRP LAESDPEAAV ELRFLASRSK AEVVDMLRKH TYLEELAQNK RAFAAISPPP 180 PKNPASSPPS SSSAAAAREH LFDKTVTPSD VGKLNRLVIP KQHAEKHFPL QIPPPTTSVV 240 ADAGAGAGEC KGVLLNFEDA AGKVWRFRYS YWNSSQSYVL TKGWSRFVKE KGLHAGDAVG 300 FYRAAGKHAQ LFIDCKVRAK STTTAAAAFL SAVAAAAAPA PAVKAIRLFG VDLLTAAAAP 360 ERRDAAGGAM TKSKRGMDAI AESQPHVVFK KQCIELALT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 3e-50 | 199 | 327 | 11 | 126 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
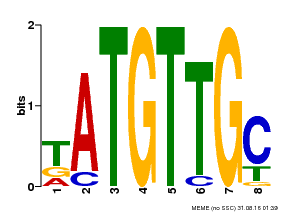
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC130725 | 0.0 | AC130725.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0494H05, complete sequence. | |||
| GenBank | AC135925 | 0.0 | AC135925.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0560C03, complete sequence. | |||
| GenBank | AC136492 | 0.0 | AC136492.3 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0554F08, complete sequence. | |||
| GenBank | AP014961 | 0.0 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012613 | 0.0 | CP012613.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015637817.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| Swissprot | Q6L4H4 | 0.0 | Y5498_ORYSJ; AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| TrEMBL | A0A0E0L5C8 | 0.0 | A0A0E0L5C8_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC05G22370.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 5e-98 | related to ABI3/VP1 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



