 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC04G26100.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 1034aa MW: 114374 Da PI: 7.3065 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.7 | 4.7e-23 | 222 | 323 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k lt sd++++g +++p++ ae+ ++++++ ++l+++d++ + W++++iyr++++r++lt+GW+ Fv a++Lk+gD+v+F +++
OPUNC04G26100.1 222 FCKNLTASDTSTHGGFSVPRRAAEKLfpqldySMQPPN-QELIVRDLHDNMWTFRHIYRGQPKRHLLTTGWSLFVGAKRLKAGDSVLFI--RDE 312
89*********************999*****9555555.49************************************************..456 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
+ +l+++v+r+
OPUNC04G26100.1 313 KSQLLLGVRRA 323
77789999997 PP
| |||||||
| 2 | Auxin_resp | 117.3 | 1.2e-38 | 348 | 430 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++s F+++YnPr+s+s Fv++v++++ka++++ svGmRf m+fete+ss+rr++Gtvvg+sd+dp+rWpnSkWr+L+
OPUNC04G26100.1 348 AAHAASSGSSFTIYYNPRTSPSPFVIPVARYNKATYMQPSVGMRFAMMFETEESSKRRYTGTVVGISDYDPMRWPNSKWRNLQ 430
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 4.19E-45 | 212 | 351 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.1E-40 | 215 | 336 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.03E-20 | 221 | 322 | No hit | No description |
| SMART | SM01019 | 1.3E-22 | 222 | 324 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.689 | 222 | 324 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.2E-20 | 222 | 323 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.3E-34 | 348 | 430 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.3E-10 | 881 | 1018 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 8.5E-9 | 928 | 1010 | No hit | No description |
| PROSITE profile | PS51745 | 25.501 | 931 | 1015 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1034 aa Download sequence Send to blast |
MRNHGFSPRK VGIFRPKQPR SAASHCSSRR ERLRSQQRER EEAKGAKGRG STTTLLRIIA 60 IYYCPHPTTP IRHHCIPPPQ PLLLPPREKA KTGVLRNAAA LLDEMQLMGE TQGAKKVINS 120 ELWHACAGPL VCLPQRGSLV YYFPQGHSEQ VAATTRKIPN SRIPNYPNLP SQLLCQVHNI 180 TLHADKDTDE VYAQMTLQPE TDVFPIPTLG AYTKSKHPTE YFCKNLTASD TSTHGGFSVP 240 RRAAEKLFPQ LDYSMQPPNQ ELIVRDLHDN MWTFRHIYRG QPKRHLLTTG WSLFVGAKRL 300 KAGDSVLFIR DEKSQLLLGV RRATRQQTTL SSSVLSTDSM HIGVLAAAAH AASSGSSFTI 360 YYNPRTSPSP FVIPVARYNK ATYMQPSVGM RFAMMFETEE SSKRRYTGTV VGISDYDPMR 420 WPNSKWRNLQ VEWDEHGYGE RPERVSIWDI ETPENTLVFP SSTLNSKRQC LPGYGVSVPG 480 LEIGSANMSS FPRAQGNPYG SLQHIPAVGS ELAIMLLNQS GQTLGSPLSF HQSSYSSIIQ 540 NVKQNYIPPL TVSTSACSTK QESMPSDDAQ HQFHMANMQN GDLESSEVQP VIDSISDSKL 600 NVTSRDPRNT DSYPIRSISE QNSKGEPRGK TRRSKKGLSH KAVSEKSDLS SAPSWICDNQ 660 QVGLESKLVG CDEQVNCGNI EDSSGALTQG NFAGQPHGHQ VEQNGVLSPP KVESSKSPDG 720 GKSVNSFPNQ VCFSQFIDGL DWMTQPSYYQ DSNVIQPADA SENIFSSSAD IPPPMIADTM 780 ETFQASCLSD CLPNSIQEFI SSPDLNSLTF LSPEMQNLEV QLQHDGSNLP STSNSFVQMS 840 FSEESGNQSA NLSGLHMESA HRSMNTTSCS QPMSTGGFDA GMHSKLPRLK ESQIMSLPEI 900 HTNSMGTSAC SMGATEYSLD RSAKPMKPPV RTYTKVQKQG SVGRSIDVTG FRNYHELRSA 960 IACMFGLQGK LEHPGSSEWK LVYVDYENDV LLVGDDPWEE FINCVRCIRI LSPSEVQQMS 1020 ENGMHVLNDC IQAS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 88 | 453 | 10 | 390 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
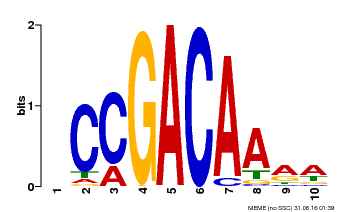
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB071292 | 0.0 | AB071292.1 Oryza sativa OsMP mRNA for Arabidopsis Monopteros-like protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025881052.1 | 0.0 | auxin response factor 11-like isoform X1 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A0A0E0KWE2 | 0.0 | A0A0E0KWE2_ORYPU; Auxin response factor | ||||
| STRING | OPUNC04G26100.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



