 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC03G37170.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 970aa MW: 105627 Da PI: 5.2219 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130 | 8.5e-41 | 152 | 229 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC+adl+ +++yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
OPUNC03G37170.1 152 ACQVEGCTADLTGVRDYHRRHKVCEMHAKATTAVVGNTVQRFCQQCSRFHPLQEFDEGKRSCRRRLAGHNRRRRKTRP 229
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.3E-34 | 145 | 214 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.394 | 150 | 227 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-38 | 151 | 231 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.6E-30 | 153 | 226 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.71E-8 | 752 | 857 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.4E-8 | 754 | 861 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.85E-9 | 757 | 857 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 970 aa Download sequence Send to blast |
MEAARVGAQS RHLYGGGGLG EPDMDRRDKR LFGWDLNDWR WDSDHFVATP VPAAEASGLV 60 LNSSPSSSEE AGAAVVRNVN VRGDSDKRKR VVVIDDDDVE DDELVENGGG SLSLRIGGDA 120 VAHGAGVGGG ADEEDRNGKK IRVQGGSSSG PACQVEGCTA DLTGVRDYHR RHKVCEMHAK 180 ATTAVVGNTV QRFCQQCSRF HPLQEFDEGK RSCRRRLAGH NRRRRKTRPE VAVGGSAFTE 240 DKVSSYLLLG LLGVCANLNA DNAEHLRGQE LLSSLLRNLG AVAKSLDPKE LCKLLEACQS 300 MQDGSNAGTS ETANALVNTA VAEAAGPSNS KMPFVNGDQC GLASSSVVPV QSKSPTVATP 360 EPPPCKFKDF DLNDTCGGME GFEDGYEGSP TPAFKTTDSP NCPSWMHQDS TQSPPQTSGN 420 SDSTSAQSLS SSNGDAQCRT DKIVFKLFEK VPSDLPPVLR SQILGWLSSS PTDIESYIRP 480 GCIILTVYLR LVESAWKELS DNMSSYLDKL LNSSTGNFWA SGLVFVMVRH QIAFMHNGQL 540 MLDRPLANSA HHYCKILCVR PIAAPFSTKV NFRVEGLNLV SDSSRLICSF EGSCIFQEDT 600 DNIVDDAEHD DIEYLNFYCP LPGSRGRGFV EVEDGGFSNG FFPFIIAEQD ICSEVCELES 660 IFESSSHEQA DDDNARNQAL EFLNELGWLL HRANVISKQD KVPLASFNLW RFRNLGIFAM 720 EREWCAVTKV LLDFLFIGLV DIGSQSPEEV VLSENLLHAA VRMKSAPMVR FLLGYKPNES 780 LKGTAETFLF RPDAQGPSKF TPLHIAAATD DAEDVLDALT NDPGLVGINA WRNARDDAGF 840 TPEDYARQRG NDAYLNMVEK KINKHLGKGH VVLGVPSSMH PVITDGAKPG EVSLEIGMSV 900 PPAAPRCNAC SRQALMYPNS TVRTFLYRPA MLTVMGIAVI CVCVGLLLHT CPKVYAAPTF 960 RWELLERGPM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-31 | 146 | 226 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
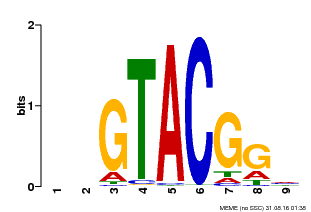
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK069847 | 0.0 | AK069847.1 Oryza sativa Japonica Group cDNA clone:J023034C10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631511.1 | 0.0 | squamosa promoter-binding-like protein 6 isoform X1 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A0E0KL59 | 0.0 | A0A0E0KL59_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC03G37130.1 | 0.0 | (Oryza punctata) | ||||
| STRING | OPUNC03G37170.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



