 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC01G12460.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 296aa MW: 32438.6 Da PI: 5.717 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.8 | 2e-09 | 230 | 263 | 22 | 55 |
SSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 22 nrypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++e++ +LA+++gL+ +q+ +WF N+R ++
OPUNC01G12460.1 230 WPYPTEEDKLRLAARTGLDPKQINNWFINQRKRH 263
59*****************************885 PP
| |||||||
| 2 | ELK | 33.2 | 1.2e-11 | 183 | 204 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsg+L+ L++EF+
OPUNC01G12460.1 183 ELKEMLLKKYSGCLSRLRSEFL 204
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 1.5E-21 | 37 | 81 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 8.6E-21 | 39 | 79 | IPR005540 | KNOX1 |
| SMART | SM01256 | 4.4E-27 | 87 | 138 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 2.7E-24 | 91 | 137 | IPR005541 | KNOX2 |
| SMART | SM01188 | 3.6E-6 | 183 | 204 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 10.423 | 183 | 203 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.4E-8 | 183 | 204 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.843 | 203 | 266 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-13 | 205 | 270 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.15E-19 | 205 | 268 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.3E-27 | 208 | 266 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.14E-13 | 215 | 267 | No hit | No description |
| Pfam | PF05920 | 6.2E-17 | 223 | 262 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 241 | 264 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010073 | Biological Process | meristem maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 296 aa Download sequence Send to blast |
MEDLYSIHPG ISRVGGAASE ASGVGVGGIG GSSDLTELMK AQIAGHPRYP TLLSAYIECR 60 KVGAPPEVAS LLEEIGRERR AGVGGGCAGE IGVDPELDEF MEAYCRVLVR YKEELSRPFD 120 EAASFLSSIQ MQLSNLCSGA TSPPATATHS DEMVGSSDED QCSGETDMLD IGQEQSSRLA 180 DHELKEMLLK KYSGCLSRLR SEFLKKRKKG KLPKDARSAL LEWWNTHYRW PYPTEEDKLR 240 LAARTGLDPK QINNWFINQR KRHWKPSDGM RFALMEGVAG GSSGTTLYFD TGTIGP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 198 | 207 | LRSEFLKKRK |
| 2 | 204 | 208 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
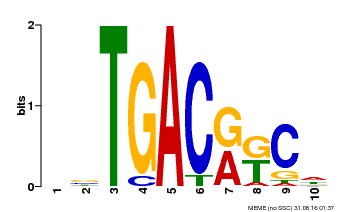
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB028883 | 0.0 | AB028883.1 Oryza sativa mRNA for knotted1-type homeobox protein OSH6, complete cds. | |||
| GenBank | AK241312 | 0.0 | AK241312.1 Oryza sativa Japonica Group cDNA, clone: J065141L09, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006644109.1 | 0.0 | PREDICTED: homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 0.0 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | A0A0E0JHJ6 | 0.0 | A0A0E0JHJ6_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC01G12460.2 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.1 | 1e-80 | KNOTTED1-like homeobox gene 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



