 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ONIVA06G28270.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 75993.4 Da PI: 7.7407 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.9 | 4.4e-24 | 128 | 229 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g +W++++iyr++++r++lt+GW+ Fv++++L +gD++vF+ ++
ONIVA06G28270.3 128 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TVLAKDVHGVVWKFRHIYRGTPRRHLLTTGWSTFVNQKKLVAGDSIVFM--RT 217
89****************************996555555..8999*********************************************..45 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
+++ l+v+++r+
ONIVA06G28270.3 218 ENGDLCVGIRRA 229
88999****997 PP
| |||||||
| 2 | Auxin_resp | 112.9 | 3.1e-37 | 293 | 376 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a ++++FevvY+Prast+eF+vk+ v++a+++++ +GmRfkmafetedss++++ +Gtv++v+ +dp+rWpnS+Wr+L+
ONIVA06G28270.3 293 AANLAVSGQPFEVVYYPRASTPEFCVKAGAVRAAMRTQWFAGMRFKMAFETEDSSRISWfMGTVSAVQVADPIRWPNSPWRLLQ 376
688999****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.9E-37 | 119 | 231 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.18E-28 | 124 | 230 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 9.50E-21 | 127 | 228 | No hit | No description |
| PROSITE profile | PS50863 | 12.999 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.4E-22 | 128 | 229 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.4E-21 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.1E-32 | 293 | 376 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 24.237 | 615 | 698 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MITFVDSAAK ERERESDKCL DPQLWHACAG GMVQMPPVSS KVYYFPQGHA EHAQGHGPVE 60 FPGGRVPALV LCRVAGVRFM ADPDTDEVFA KIRLVPVRAN EQGYAGDADD GIGAAAAAAA 120 QEEKPASFAK TLTQSDANNG GGFSVPRYCA ETIFPRLDYS ADPPVQTVLA KDVHGVVWKF 180 RHIYRGTPRR HLLTTGWSTF VNQKKLVAGD SIVFMRTENG DLCVGIRRAK KGGVGGPEFL 240 PPPPPPPPPT PAAGGNYGGF SMFLRGDDDG NKMAAAARGK VRARVRPEEV VEAANLAVSG 300 QPFEVVYYPR ASTPEFCVKA GAVRAAMRTQ WFAGMRFKMA FETEDSSRIS WFMGTVSAVQ 360 VADPIRWPNS PWRLLQVSWD EPDLLQNVKR VSPWLVELVS NMPAIHLAPF SPPRKKLCVP 420 LYPELPIDGQ FPTPMFHGNP LARGVGPMCY FPDGTPAGIQ GARHAQFGIS LSDLHLNKLQ 480 SSLSPHGLHQ LDHGMQPRIA AGLIIGHPAA RDDISCLLTI GSPQNNKKSD AKKAPAQLML 540 FGKPILTEQQ ISLGDAASVA VKKSSSDGNA ENTVNKSNSD VSSPRSNQNG TTDNLSCGGV 600 PLCQDNKVLD VALETGHCKV FMQSEDVGRT LDLSVVGSYE ELYRRLADMF SIEKAELMSH 660 VFYRDAAGAL KHTGDEPFSE FTKTARRLNI LTDTSGDNLA R |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 5e-78 | 20 | 397 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
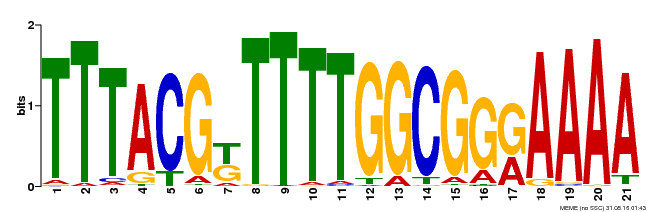
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK100322 | 0.0 | AK100322.1 Oryza sativa Japonica Group cDNA clone:J023079H12, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015644137.1 | 0.0 | auxin response factor 18 | ||||
| Refseq | XP_015644138.1 | 0.0 | auxin response factor 18 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A0E0HUP0 | 0.0 | A0A0E0HUP0_ORYNI; Auxin response factor | ||||
| STRING | ONIVA06G28270.1 | 0.0 | (Oryza nivara) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



