 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ONIVA04G27140.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 1039aa MW: 114354 Da PI: 6.8923 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.7 | 4.7e-23 | 227 | 328 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k lt sd++++g +++p++ ae+ ++++++ ++l+++d++ + W++++iyr++++r++lt+GW+ Fv a++Lk+gD+v+F +++
ONIVA04G27140.1 227 FCKNLTASDTSTHGGFSVPRRAAEKLfpqldySMQPPN-QELIVRDLHDNMWTFRHIYRGQPKRHLLTTGWSLFVGAKRLKAGDSVLFI--RDE 317
89*********************999*****9555555.49************************************************..456 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
+ +l+++v+r+
ONIVA04G27140.1 318 KSQLLLGVRRA 328
77789999997 PP
| |||||||
| 2 | Auxin_resp | 117.3 | 1.3e-38 | 353 | 435 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++s F+++YnPr+s+s Fv++v++++ka++++ svGmRf m+fete+ss+rr++Gtvvg+sd+dp+rWpnSkWr+L+
ONIVA04G27140.1 353 AAHAASSGSSFTIYYNPRTSPSPFVIPVARYNKATYMQPSVGMRFAMMFETEESSKRRYTGTVVGISDYDPMRWPNSKWRNLQ 435
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.92E-45 | 217 | 356 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 4.8E-40 | 220 | 341 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.05E-20 | 226 | 327 | No hit | No description |
| SMART | SM01019 | 1.3E-22 | 227 | 329 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.689 | 227 | 329 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.3E-20 | 227 | 328 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.3E-34 | 353 | 435 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.3E-10 | 890 | 1023 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 8.5E-9 | 933 | 1015 | No hit | No description |
| PROSITE profile | PS51745 | 25.501 | 936 | 1020 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1039 aa Download sequence Send to blast |
MRNHGFPPRK VGIFRAKQPR SAASHTGAAA AAAGERLGSQ DQEREGKRRK GDEGSTTTLL 60 RIIAIYCPHP TTTPTPVYST PQPLLLPPRE KAKTGVLRNA AALLDEMQLM GETQGAKKVI 120 NSELWHACAG PLVCLPQRGS LVYYFPQGHS EQVAATTRKI PNSRIPNYPN LPSQLLCQVH 180 NITLHADKDT DEVYAQMTLQ PVNSETDVFP IPTLGAYTKS KHPTEYFCKN LTASDTSTHG 240 GFSVPRRAAE KLFPQLDYSM QPPNQELIVR DLHDNMWTFR HIYRGQPKRH LLTTGWSLFV 300 GAKRLKAGDS VLFIRDEKSQ LLLGVRRATR QQTMLSSSVL STDSMHIGVL AAAAHAASSG 360 SSFTIYYNPR TSPSPFVIPV ARYNKATYMQ PSVGMRFAMM FETEESSKRR YTGTVVGISD 420 YDPMRWPNSK WRNLQVEWDE HGYGERPERV SIWDIETPEN TLVFPSSTLN SKRQCLPGYG 480 VSVPGMEIGS ANMSSFPRAQ GNPYGSLQHI PAVGSELAIM LLNQSGQTLG SPLSFHQSSY 540 SSIIQNVKQN YIPPLTVSTS ACLTKQESLP SDDAQHQFHM ANMQNGDLEG SEVQPVIDSI 600 SESKLNATSR DPRNTDPYTS RSTSEQNSKG EPRGKTRRSK KGLPHKTVSE KSDLSSAPSW 660 ICDNQQVGLE SKLVGCDEQV NCGNIEDSSG ALTQGNFVGQ PHGHQVEQKG VLSPPKVESS 720 KSPDGGKSVN SFPNQGCFSQ FIDGLDWMTQ PSYYQDSNVI QPAGVSENIF SSSADIPPSM 780 IADTMETFQA SCLSDCLPNS IQEFISSPDL NSLTFLSPDM QNLEVQLQHD GSNLPSTSNS 840 FVQMSFSEES ASQSANLSGL HMESTHRSIN TTSCSQPMST GGFDAGMYSK LPRLKESQIL 900 SLPEIHTNSM GTSACSMDAT EYSLDRSAKP MKPPVRTYTK VQKQGSVGRS IDVTGFRNYH 960 ELRSAIACMF GLQGKLEHPG SSEWKLVYVD YENDVLLVGD DPWEEFINCV RCIRILSPSE 1020 VQQMSENGMH VLNDCIQAA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 90 | 458 | 10 | 390 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
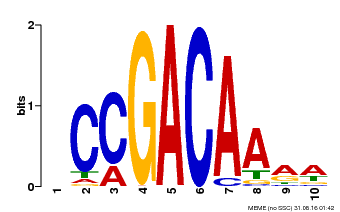
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB071292 | 0.0 | AB071292.1 Oryza sativa OsMP mRNA for Arabidopsis Monopteros-like protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025881052.1 | 0.0 | auxin response factor 11-like isoform X1 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A0A0E0H738 | 0.0 | A0A0E0H738_ORYNI; Auxin response factor | ||||
| STRING | ONIVA04G27140.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



