 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ONIVA01G43630.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 636aa MW: 69115.8 Da PI: 6.7413 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 19.4 | 2.8e-06 | 397 | 418 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C +Cgk F+r nL+ H+r H
ONIVA01G43630.1 397 FCLICGKGFKRDANLRMHMRGH 418
5*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 9.05E-5 | 394 | 422 | No hit | No description |
| SMART | SM00355 | 0.012 | 396 | 418 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.552 | 396 | 423 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-4 | 398 | 422 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 398 | 418 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 68 | 445 | 478 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 22 | 483 | 505 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 636 aa Download sequence Send to blast |
MENCLRSTGA RNQQGCSSYK ATHPPSRVPF LLPCGASNST RVLLHLQLHI RSDPCYLVPP 60 LSLLRNRLAV RHHGDSTSIS PRSPSTPPLA AAAALAVAVA SADPIPIHPG DREQGAGGLG 120 RSSETSLKAL PSMASNATRN TDPDQQGVRF SSMDQPPCFA RPGQSFPAFP PLFGVQSSSL 180 YLPDDIEAKI GNQFESNPSP NNPTMDWDPQ AMLSNLSFLE QKIKQVKDIV QSMSNRESQV 240 AGGSSEAQAK QQLVTADLTC IIIQLISTAG SLLPSMKNPI SSNPALRHLS NTLCAPMILG 300 TNCNLRPSAN DEATIPDISK THDYEELMNS LNTTQAESDE MMNCQNPCGG EGSEPIPMED 360 HDVKESDDGG ERENLPPGSY VVLQLEKEEI LAPHTHFCLI CGKGFKRDAN LRMHMRGHGD 420 EYKTAAALAK PSKDSSSESA PVTRYSCPYV GCKRNKEHKK FQPLKTILCV KNHYKRSHCD 480 KSYTCSRCNT KKFSVIADLK THEKHCGRDK WLCSCGTTFS RKDKLFGHVA LFQGHTPALP 540 MDDIKVTGAS EQPQGSEAMN TMVGSAGYNF PGSSSDDIPN LDMKMADDPR YFSPLSFDPC 600 FGGLDDFTRP GFDISENPFS FLPSGSCSFG QQNGDS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in aluminum tolerance. {ECO:0000250}. | |||||
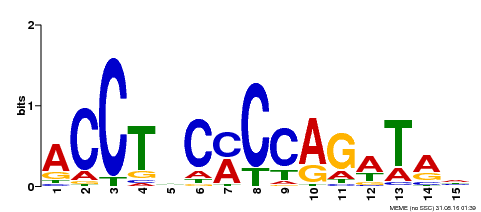
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012609 | 0.0 | CP012609.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015641300.1 | 0.0 | zinc finger protein STOP1 homolog | ||||
| Refseq | XP_015641307.1 | 0.0 | zinc finger protein STOP1 homolog | ||||
| Swissprot | Q943I6 | 0.0 | STOP1_ORYSJ; Zinc finger protein STOP1 homolog | ||||
| TrEMBL | A0A0E0FWB8 | 0.0 | A0A0E0FWB8_ORYNI; Uncharacterized protein | ||||
| STRING | ONIVA01G43630.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1512 | 38 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 1e-156 | C2H2 family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



