 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI09G11520.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 378aa MW: 39212.2 Da PI: 8.4246 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.3 | 1.4e-40 | 60 | 133 | 4 | 77 |
STT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 4 vegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
vegC +dls +k y+ rhkvC +h+k p v+v+gleqrfCqqCsrfh+l efD+ek+sCrrrLa+hnerrrk++
OMERI09G11520.1 60 VEGCGVDLSGVKPYYCRHKVCYMHAKEPIVVVAGLEQRFCQQCSRFHQLPEFDQEKKSCRRRLAGHNERRRKPT 133
89**********************************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51141 | 29.145 | 55 | 132 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.5E-31 | 60 | 131 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.57E-37 | 60 | 135 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:4.10.1100.10 | 1.1E-29 | 60 | 119 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:2000025 | Biological Process | regulation of leaf formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 378 aa Download sequence Send to blast |
MATGGSGGGG GGGGGGGDDV HGLKFGKKIY FEQDAAAAVE SSSTSSGGGG GGGGKKGKGV 60 EGCGVDLSGV KPYYCRHKVC YMHAKEPIVV VAGLEQRFCQ QCSRFHQLPE FDQEKKSCRR 120 RLAGHNERRR KPTPGPLSSR YGRLAASFHE EPGRSRSFVV DFSYPRVPSS VRDAWPAIQP 180 GDRMSGSIQW QGGHELHPHR SSAVAGYGDH HAFSSHGGSA AGAPMLHHPA FELTSGGCLA 240 GVATDSSCAL SLLSTQPWDT TQSTSSHNRS PPMSSTASAF GGGNNPVSPS VMASNYMAAS 300 PGWNSSSRGH DGARNVHLPP PHGVALNEVP PGSVHHGHFS GELELALQGG ALSNRPEAEH 360 GSGSGAFSHS TNAMNWSL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-27 | 48 | 131 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00307 | DAP | Transfer from AT2G42200 | Download |
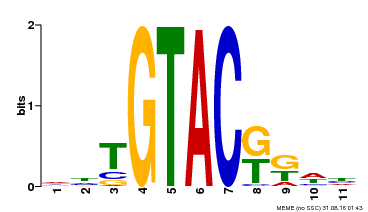
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK063561 | 0.0 | AK063561.1 Oryza sativa Japonica Group cDNA clone:001-117-F03, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015610961.1 | 0.0 | squamosa promoter-binding-like protein 17 isoform X1 | ||||
| Swissprot | A3C057 | 0.0 | SPL17_ORYSJ; Squamosa promoter-binding-like protein 17 | ||||
| TrEMBL | A0A0E0ETJ4 | 0.0 | A0A0E0ETJ4_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI09G11350.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2527 | 37 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42200.1 | 5e-43 | squamosa promoter binding protein-like 9 | ||||



