 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI06G27280.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 237aa MW: 24151.8 Da PI: 8.4138 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 218.5 | 1.5e-67 | 12 | 164 | 4 | 154 |
DUF702 4 gtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasa.....aeaaskrkrel..kskkqsalsstk 90
++sCqdCGn+akkdC+h RCRtCC+srgf+CathvkstWvpaakrrerqqqlaa+ + aaa++ a+aaskr+rel ++++a+ +t+
OMERI06G27280.1 12 AGTSCQDCGNNAKKDCSHLRCRTCCRSRGFSCATHVKSTWVPAAKRRERQQQLAALFRGAAANNsaaaaAAAASKRPRELvrLPSANTAMVATT 105
4579***************************************************999998888777777788889998733344444444444 PP
DUF702 91 lssaeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGlee 154
+ss+ ++P+e+s+eavfrcvr+++vd++++elaYqtavsigGh+fkGiL+d+G+++
OMERI06G27280.1 106 TSSG-----EGDGRFPPELSVEAVFRCVRIGAVDEADAELAYQTAVSIGGHTFKGILRDHGPAD 164
4444.....45678***********************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 8.0E-64 | 13 | 163 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 1.0E-24 | 15 | 57 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 4.1E-20 | 114 | 162 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010252 | Biological Process | auxin homeostasis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0048480 | Biological Process | stigma development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 237 aa Download sequence Send to blast |
MGGPGGAGGG RAGTSCQDCG NNAKKDCSHL RCRTCCRSRG FSCATHVKST WVPAAKRRER 60 QQQLAALFRG AAANNSAAAA AAAASKRPRE LVRLPSANTA MVATTTSSGE GDGRFPPELS 120 VEAVFRCVRI GAVDEADAEL AYQTAVSIGG HTFKGILRDH GPADEAAGQL PPSSAEYHQL 180 TGQGREESSP AGSSEGVGGG HGAATAATSA AVLMDPYPTP IGAFAAGTQF FPHNPRT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). {ECO:0000269|PubMed:12361963, ECO:0000269|PubMed:16740145, ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:18811619, ECO:0000269|PubMed:20154152, ECO:0000269|PubMed:22318676}. | |||||
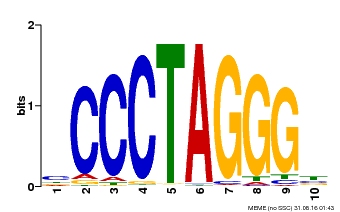
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00613 | PBM | Transfer from AT3G51060 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Regulated by ESR1 and ESR2. {ECO:0000269|PubMed:21976484}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004329 | 0.0 | AP004329.3 Oryza sativa Japonica Group genomic DNA, chromosome 6, BAC clone:OJ1663_H12. | |||
| GenBank | AP005463 | 0.0 | AP005463.3 Oryza sativa Japonica Group genomic DNA, chromosome 6, PAC clone:P0712G01. | |||
| GenBank | AP014962 | 0.0 | AP014962.1 Oryza sativa Japonica Group DNA, chromosome 6, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012614 | 0.0 | CP012614.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 6 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015642175.1 | 1e-135 | protein SHI RELATED SEQUENCE 1 | ||||
| Refseq | XP_025881792.1 | 1e-135 | protein SHI RELATED SEQUENCE 1 | ||||
| Swissprot | Q9SD40 | 4e-62 | SRS1_ARATH; Protein SHI RELATED SEQUENCE 1 | ||||
| TrEMBL | A0A0E0E6A5 | 1e-171 | A0A0E0E6A5_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI06G25600.1 | 1e-171 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1354 | 35 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G51060.1 | 1e-56 | Lateral root primordium (LRP) protein-related | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



