 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI06G15340.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 458aa MW: 48899.1 Da PI: 4.6543 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 87.6 | 1.2e-27 | 183 | 237 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k++++WtpeLH+rFv+aveqL G++kA+P++ile+m++++Lt+++++SHLQkYR++
OMERI06G15340.1 183 KAKVDWTPELHRRFVQAVEQL-GIDKAVPSRILEIMGIDSLTRHNIASHLQKYRSH 237
6799*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.924 | 180 | 239 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.63E-19 | 181 | 240 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-27 | 181 | 240 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.7E-26 | 183 | 237 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.3E-8 | 186 | 235 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 458 aa Download sequence Send to blast |
MLAVSPAMCP DIEERAAVAG DDGMEVVGMS SSDMDQFDFS VDDIDFGDFF LRLEDGDVLP 60 DLEVDPAEIF TDFEAIATSG GEGVQDQEVP TVELLAPADD VGVLDPCGDV VVGEENAAFA 120 EAGEEKGGCN QDDDAGEANV DDGADGAAAV EAKSSSPSST TSSSQEAESR HKSSSKSSHG 180 KKKAKVDWTP ELHRRFVQAV EQLGIDKAVP SRILEIMGID SLTRHNIASH LQKYRSHRKH 240 MIAREAEAAS WTQRRHIYAA GGGAVAKRPE SNAWAVPTIG FPPPPPPPPS PASMQHYARP 300 LHVWGHPTMD LSRVPVWPPR HLVPRGPAPP WVPPPPPSDP VLWHHPYMRG PAHVPTQGTP 360 CMAMPMPAAR FPAPPVPGVV PCPMYRPLTP PALAIKNQQD AQLQLQVQPS SESIDAAIGD 420 VLSKPWLPLP LGLKPPSVDS VMGELQRQGV ANVPPACG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 5e-17 | 180 | 235 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport. {ECO:0000269|PubMed:19808806}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
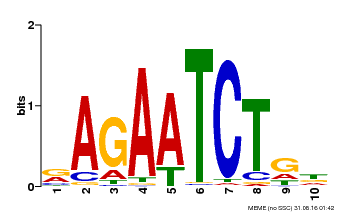
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK061946 | 0.0 | AK061946.1 Oryza sativa Japonica Group cDNA clone:001-042-E04, full insert sequence. | |||
| GenBank | AK065184 | 0.0 | AK065184.1 Oryza sativa Japonica Group cDNA clone:J013002D22, full insert sequence. | |||
| GenBank | AK098909 | 0.0 | AK098909.1 Oryza sativa Japonica Group cDNA clone:J013002F22, full insert sequence. | |||
| GenBank | AK122161 | 0.0 | AK122161.1 Oryza sativa Japonica Group cDNA clone:J033147G01, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015644244.1 | 0.0 | probable transcription factor GLK1 | ||||
| Swissprot | Q5Z5I4 | 0.0 | GLK1_ORYSJ; Probable transcription factor GLK1 | ||||
| TrEMBL | A0A0E0E1L0 | 0.0 | A0A0E0E1L0_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI06G14290.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2415 | 38 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 2e-49 | GBF's pro-rich region-interacting factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



