 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI05G16220.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 318aa MW: 34352.6 Da PI: 7.0058 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.2 | 7.6e-08 | 42 | 86 | 2 | 44 |
SSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrw 44
g+W++eE + + +a + l + +W+++a ++ g+t ++ ++
OMERI05G16220.1 42 GAWSQEENKVFEQALAALDRNdpeRWERVALLLP-GKTVADVMTHY 86
79*****************99*************.********999 PP
| |||||||
| 2 | Myb_DNA-binding | 45.6 | 1.6e-14 | 156 | 200 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE++l++ + k++G g+W+ I+r + + Rt+ q+ s+ qky
OMERI05G16220.1 156 PWTEEEHKLFLMGLKKYGRGDWRNISRNFVTSRTPTQVASHAQKY 200
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 8.8E-7 | 40 | 92 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.8E-5 | 42 | 86 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 2.3E-9 | 42 | 97 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 6.249 | 42 | 90 | IPR017877 | Myb-like domain |
| CDD | cd00167 | 5.09E-6 | 43 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 17.568 | 149 | 205 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.44E-17 | 151 | 205 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.0E-17 | 152 | 203 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.8E-13 | 153 | 203 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-12 | 153 | 199 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 7.5E-12 | 156 | 200 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.18E-11 | 156 | 201 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 318 aa Download sequence Send to blast |
MMKESYMEVL PPAPAHYFVG QAAAAAAAGG WFLPDRRGGG GGAWSQEENK VFEQALAALD 60 RNDPERWERV ALLLPGKTVA DVMTHYDDLE NDVCFIEAGL VPFPHYGGGG SPSSAGFTLD 120 WDGGDDPAGL GFKRSCYMVG GKRARGPDQE RKKGVPWTEE EHKLFLMGLK KYGRGDWRNI 180 SRNFVTSRTP TQVASHAQKY FIRLNSGGKD KRRSSIHDIT TVNLPDDDHG NPSPSPPPSV 240 LTAHSSSSAT AVSEQFGVLV DGKPPPPLGR GAGHHHFMPH PYAQVKIEAG NSHVAGGGRL 300 DDSVLMQMQC GQLMQPLG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 2e-15 | 39 | 110 | 6 | 77 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
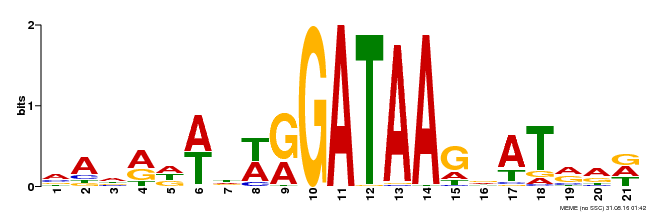
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK068138 | 0.0 | AK068138.1 Oryza sativa Japonica Group cDNA clone:J013135D01, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015637475.1 | 0.0 | transcription factor DIVARICATA | ||||
| Swissprot | Q8S9H7 | 4e-81 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A0E0DS94 | 0.0 | A0A0E0DS94_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI05G15400.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2274 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 4e-81 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



