 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI03G06500.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 989aa MW: 110750 Da PI: 5.7335 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.8 | 2.8e-55 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
ke ++rwl+++ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een +fq
OMERI03G06500.1 20 KEaQRRWLRPTEICEILKNYRSFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENINFQ 113
55599***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Lee++ +ivlvhylevk
OMERI03G06500.1 114 RRSYWMLEEDYMHIVLVHYLEVK 136
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.727 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 5.2E-76 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.5E-48 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.1E-7 | 401 | 487 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 3.8E-8 | 402 | 480 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 6.82E-18 | 402 | 488 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00102 | 5.68E-4 | 402 | 489 | No hit | No description |
| SuperFamily | SSF48403 | 9.95E-17 | 591 | 695 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.3E-18 | 593 | 699 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.00E-13 | 594 | 692 | No hit | No description |
| PROSITE profile | PS50297 | 18.059 | 600 | 704 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.763 | 600 | 632 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.013 | 633 | 662 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.843 | 633 | 665 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1400 | 672 | 701 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 4.51E-8 | 804 | 857 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.075 | 806 | 828 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.407 | 807 | 836 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0032 | 808 | 827 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 829 | 851 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 830 | 854 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.9E-4 | 832 | 851 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 989 aa Download sequence Send to blast |
MTEGRRYAIA PQLDIEQILK EAQRRWLRPT EICEILKNYR SFRIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKSGSIDVLH CYYAHGEENI NFQRRSYWML 120 EEDYMHIVLV HYLEVKAGKL SSRSTGHDDV LQASHADSPL SQLPSQTTEG ESSVSGQASE 180 YDETESGSYQ GLQATAPNTG FYSHGQDNLP VVLNESDLGT AFNGPNSQFD LSLWIEAMRP 240 DKGTHQIPLY QAPVPSEQSP FTGGPGIESF TFDEVYNNGL SIKDVDGDDT DGETPWQIPN 300 ASGTFATVDS FQQNDKTLEE AINYPLLKTQ SSSLSDIIKD SFKKNDSFTR WMSKELAEVD 360 DSQITSSSGV YWNSEEADSI IEASSSDQYT LGPVLAQDQL FTIVDFSPTW TYAGSKTRVF 420 IKGNFLSSDE VKRLKWSCMF GEFEVPAEII ADDTLVCHSP SHKPGRVPFY VTCSNRLACS 480 EVREFEFRPQ YMDAPSPLGS TNKIYLQKRL DKLLSLEQDE IQTTLSNPTN EIIDLSKKIS 540 SLMMNNDDWS ELLKLADDNE PATDDKQDQF LQNRIKEKLH IWLLHKVGDG GKGPSVLDEE 600 GQGVLHLAAA LGYDWAIRPT IAAGVNINFR DAHGWTALHW AAFCGRERTV VALIALGAAP 660 GAVTDPTPSF PSGSTPADLA SANGHKGISG FLAESSLTSH LQTLNLKEAM RSSAGEISGL 720 PGIVNVADRS ASPLAVEGHQ TGSMGDSLGA VRNAAQAAAR IYQVFRMQSF QRKQAVQYED 780 ENGAISDERA MSLLSAKPSK PAQLDPLHAA ATRIQNKFRG WKGRKEFLLI RQRIVKIQAH 840 VRGHQVRKHY RKIIWSVGIV EKVILRWRRR GAGLRGFRPT ENAVTESTSS SSGNVTQNRP 900 AENDYDFLQE GRKQTEERLQ KALARVKSMV QYPDARDQYQ RILTVVTKMQ ESQAMQEKML 960 EESTEMDEGL LMSEFKELWD DDMPTPGYF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
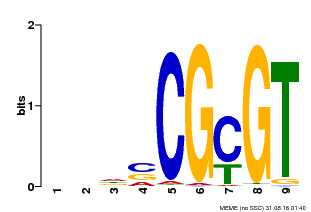
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK066651 | 0.0 | AK066651.1 Oryza sativa Japonica Group cDNA clone:J013074G10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631249.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A0E0CWH5 | 0.0 | A0A0E0CWH5_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI03G06390.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



