 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI02G32980.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 365aa MW: 40169.6 Da PI: 4.644 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58.2 | 2e-18 | 119 | 168 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+++G+r+++ +g+W+AeIrdp++ r++lg+f +aeeAa+a++a +++++g
OMERI02G32980.1 119 QFRGIRQRP-WGKWAAEIRDPRK---GVRVWLGTFNSAEEAARAYDAEARRIRG 168
69*******.**********954...4************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 9.9E-36 | 119 | 182 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.58E-31 | 119 | 178 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 3.6E-31 | 119 | 177 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.05E-21 | 119 | 176 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 23.459 | 119 | 176 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.6E-11 | 120 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-10 | 120 | 131 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-10 | 142 | 158 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0070483 | Biological Process | detection of hypoxia | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 365 aa Download sequence Send to blast |
MCGGAIIHHL KGHPEGSRRV TEGLLWPEKK KPRWGGGGRR HFGGFVEEDD EDFEADFEEF 60 EVDSGDSDLE LGEEDDDDVV EIKPAAFKRA LSRDNLSTIT TAGFDGPAAK SAKRKRKNQF 120 RGIRQRPWGK WAAEIRDPRK GVRVWLGTFN SAEEAARAYD AEARRIRGKK AKVNFPEAPT 180 TAQKRRAGST TAKAPKSSVE QKPTVKPAFN NLANANAFVY PSANFTSNKP FVQPDNMPFV 240 PAMNSAAPIE DPIINSDQGS NSFGCSDFGW ENDTKTPDIT SIAPISTIAE VDESAFIKSS 300 TNPMVPPVME NSAVDLPDLE PYMRFLLDDG AGDSIDSLLN LDGSQDVVSN MDLWSFDDMP 360 VSDFY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 4e-19 | 119 | 177 | 14 | 73 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that may be involved in defense signaling pathway. Binds in vitro to the DNA sequence 5'-AGCCGCC-3' of the GCC-box element found in pathogenesis-related (PR) gene promoters. The transcriptional activation is enhanced in vitro by the presence of MPK12/BWMK1. {ECO:0000269|PubMed:12913152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00204 | DAP | Transfer from AT1G53910 | Download |
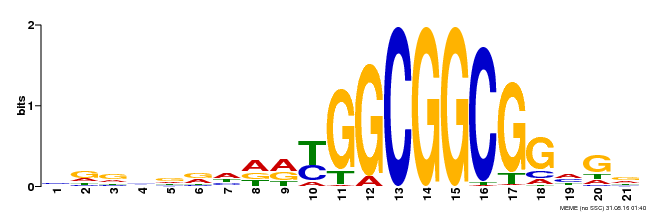
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK064027 | 0.0 | AK064027.1 Oryza sativa Japonica Group cDNA clone:001-125-B02, full insert sequence. | |||
| GenBank | AK103783 | 0.0 | AK103783.1 Oryza sativa Japonica Group cDNA clone:J033145H23, full insert sequence. | |||
| GenBank | EU837255 | 0.0 | EU837255.1 Oryza sativa Japonica Group clone KCS057H03 EREBP transcription factor mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015625463.1 | 0.0 | LOW QUALITY PROTEIN: ethylene-responsive transcription factor 1-like | ||||
| Swissprot | Q6K7E6 | 0.0 | ERF1_ORYSJ; Ethylene-responsive transcription factor 1 | ||||
| TrEMBL | A0A0E0CS91 | 0.0 | A0A0E0CS91_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI02G32230.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1039 | 38 | 134 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53910.3 | 3e-36 | related to AP2 12 | ||||



