 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI01G31990.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 530aa MW: 57280.6 Da PI: 5.0126 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.6 | 1.1e-16 | 45 | 92 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+vk++G g+W+++ + g+ R++k+c++rw ++l
OMERI01G31990.1 45 KGPWTSAEDAILVDYVKKHGEGNWNAVQKNTGLFRCGKSCRLRWANHL 92
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 51 | 3.2e-16 | 98 | 141 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l+++++ ++G++ W++ a++++ gRt++++k++w++
OMERI01G31990.1 98 KGAFTAEEERLIIQLHSKMGNK-WARMAAHLP-GRTDNEIKNYWNT 141
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.625 | 40 | 92 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-31 | 42 | 139 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-14 | 44 | 94 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.0E-14 | 45 | 92 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-23 | 46 | 99 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.49E-11 | 47 | 92 | No hit | No description |
| PROSITE profile | PS51294 | 25.026 | 93 | 147 | IPR017930 | Myb domain |
| SMART | SM00717 | 6.3E-17 | 97 | 145 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.6E-15 | 98 | 141 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-25 | 100 | 145 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.03E-12 | 100 | 141 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 530 aa Download sequence Send to blast |
MYRVKSESDC EMIHQEQMDS PVADDGSSGG SPHRGGGGGG PPLKKGPWTS AEDAILVDYV 60 KKHGEGNWNA VQKNTGLFRC GKSCRLRWAN HLRPNLKKGA FTAEEERLII QLHSKMGNKW 120 ARMAAHLPGR TDNEIKNYWN TRIKRCQRAG LPIYPTSVCN QSSNEDQQCS SDFDCGENLS 180 NDLLNANAVS ISNLLGQSFA SKSCSFMDQV NQTGMLKQSD GVLPGLSDTI NGVLSSVDQF 240 SNDSEKLKQA VGFDYLHEAN SSSKIIAPFG GALNGSHAFL NGNFSASRPT SGPLKMELPS 300 LQDTESDPNS WLKYTVAPAL QPTELVDPYL QSPAATPSVK SECASPRNSG LLEELIHEAQ 360 TLRSGKNQQT SVISSSSSVG TPCNTTVLSP EFDMCQEYWE EQHPGPFLND CAPFSGNSFT 420 ESTPPVSAAS PDIFQLSKVS PAQSTSMGSG EQVMGPKYEP GDTSPHPENF RPDALFSGNT 480 ADPSVFNNAI AMLLGNDLSI DCRPVLGDGI MFNSSSWSNM PHACEMSEFK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-32 | 39 | 145 | 1 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
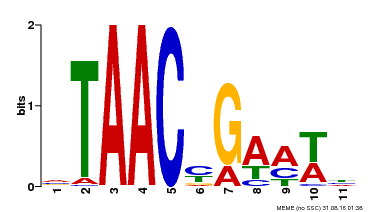
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CT830881 | 0.0 | CT830881.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCFA206M18, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015622335.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_015622336.1 | 0.0 | transcription factor GAMYB | ||||
| Swissprot | A2WW87 | 0.0 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A0E0C9C3 | 0.0 | A0A0E0C9C3_9ORYZ; Transcription factor | ||||
| STRING | OGLUM01G20920.1 | 0.0 | (Oryza glumipatula) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-88 | myb domain protein 33 | ||||



