 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 9237 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 76aa MW: 8572.5 Da PI: 11.8981 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 73.7 | 3e-23 | 19 | 69 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GVr+++ +g+W+AeIrdp + r +r +lg+f+taeeAa+a++aa++ ++g
9237 19 SQYRGVRQRP-WGSWAAEIRDP---N-RgARLWLGTFDTAEEAARAYDAAARHIRG 69
78********.**********8...3.35***********************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF038123 | 3.4E-25 | 1 | 76 | IPR017392 | AP2/ERF transcription factor ERF/PTI6 |
| Pfam | PF00847 | 2.1E-16 | 19 | 69 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.43E-16 | 19 | 76 | No hit | No description |
| SuperFamily | SSF54171 | 2.48E-23 | 19 | 76 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 23.894 | 20 | 76 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 8.0E-32 | 20 | 76 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.6E-34 | 20 | 76 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.5E-12 | 21 | 32 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.5E-12 | 43 | 59 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 76 aa Download sequence Send to blast |
RRRTQGGKGD KQSKKATTSQ YRGVRQRPWG SWAAEIRDPN RGARLWLGTF DTAEEAARAY 60 DAAARHIRGP NAKTNF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 1e-20 | 5 | 76 | 3 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that may be involved in defense signaling pathway. Binds in vitro to the DNA sequence 5'-AGCCGCC-3' of the GCC-box element found in pathogenesis-related (PR) gene promoters. The transcriptional activation is enhanced in vitro by the presence of MPK12/BWMK1. {ECO:0000269|PubMed:12913152}. | |||||
| UniProt | Transcription factor that binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. Probably acts as a transcriptional repressor and may regulate other AtERFs (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
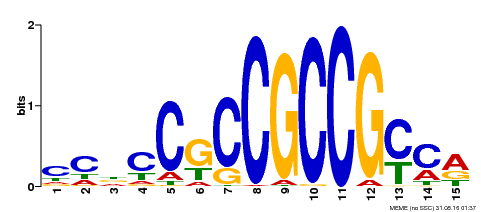
| Motif ID | Method | Source | Motif file |
| MP00289 | DAP | Transfer from AT2G33710 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Strongly and quickly induced by ethylene. {ECO:0000269|PubMed:10945353}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP000584 | 1e-124 | CP000584.1 Ostreococcus lucimarinus CCE9901 chromosome 4, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001417371.1 | 1e-48 | predicted protein, partial | ||||
| Swissprot | Q6K7E6 | 2e-25 | ERF1_ORYSJ; Ethylene-responsive transcription factor 1 | ||||
| Swissprot | Q9LW49 | 2e-25 | ERF4_NICSY; Ethylene-responsive transcription factor 4 | ||||
| TrEMBL | A4RWQ8 | 2e-47 | A4RWQ8_OSTLU; Uncharacterized protein (Fragment) | ||||
| STRING | ABO95664 | 4e-48 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP132 | 15 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G11140.1 | 5e-20 | cytokinin response factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 9237 |



