 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 25091 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 511aa MW: 54728.3 Da PI: 8.9536 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 48.2 | 2.1e-15 | 194 | 232 | 3 | 42 |
TCR 3 kkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkeek 42
+k+CnCkkskClk+YCeCfaag++C++ C+C++C+N++e+
25091 194 SKKCNCKKSKCLKLYCECFAAGAFCKD-CSCQQCQNTTEN 232
79*************************.********9875 PP
| |||||||
| 2 | TCR | 52.7 | 8.5e-17 | 264 | 303 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
+++kgC+Ckks ClkkYCeCf+ag+kC++ CkCe+CkN++
25091 264 RHTKGCHCKKSACLKKYCECFQAGVKCQDYCKCEGCKNND 303
589***********************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 1.2E-13 | 192 | 232 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 32.257 | 193 | 304 | IPR005172 | CRC domain |
| Pfam | PF03638 | 8.8E-12 | 195 | 229 | IPR005172 | CRC domain |
| SMART | SM01114 | 1.3E-14 | 264 | 305 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 1.1E-12 | 267 | 302 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 511 aa Download sequence Send to blast |
MNRVRLTNAS DDARRRSDGD GKATNGGGVS DVARLALSAE ASIQRGNGGG GRARRQLDVE 60 FGELDVQALQ DDRYVLQNMY DEFDDSVIGV EEQGSYPRRG GASRGDGHED SASKRSSTTG 120 ASVEWTKGYT KTLKNFDADS PRVGTRSARA PRGGVEASKA VVAKRGVATA KNSSLASNAS 180 AMKNASACRE FVASKKCNCK KSKCLKLYCE CFAAGAFCKD CSCQQCQNTT ENEAIVTKTR 240 QQIEQRNPYA FESKIMADAG DDARHTKGCH CKKSACLKKY CECFQAGVKC QDYCKCEGCK 300 NNDNGPSPAL PRGGAAKASK VSKSKARSRE TVAATTMAAK ELQDDLIIED FKLTGVMGSP 360 LRAFEHTDDL SSEYSNLSKS VEQSPLRTLL LSEGVQLSPF FSTSSLPATA LSPLRAGQMS 420 PLRPTPSHGF MSPLTPGMRP GKYSIRSSSS RGKAPVPLFN EDSGHGGEFK TPRDRKTNTV 480 FGGMHDAAKD GPLRATFTSP IPLSTPDVYE * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 1e-21 | 195 | 306 | 12 | 125 | Protein lin-54 homolog |
| 5fd3_B | 1e-21 | 195 | 306 | 12 | 125 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
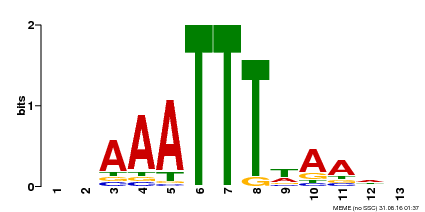
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP000589 | 0.0 | CP000589.1 Ostreococcus lucimarinus CCE9901 chromosome 9, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001419860.1 | 0.0 | predicted protein | ||||
| TrEMBL | A4S357 | 0.0 | A4S357_OSTLU; Uncharacterized protein | ||||
| STRING | ABO98153 | 0.0 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP323 | 15 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14770.1 | 8e-27 | TESMIN/TSO1-like CXC 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 25091 |



