 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN540903.1_FGP004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1026aa MW: 114661 Da PI: 6.6673 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.4 | 2.1e-56 | 21 | 137 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenpt 93
l+ ++rwl+++ei++iL n++k++++ e+++rp sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++
KN540903.1_FGP004 21 LEAQNRWLRPTEICHILSNYKKFSIAPEPPNRPASGSLFLFDRKILRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENEN 112
5679**************************************************************************************** PP
CG-1 94 fqrrcywlLeeelekivlvhylevk 118
fqrr+ywlLee + +ivlvhylevk
KN540903.1_FGP004 113 FQRRTYWLLEEGFMNIVLVHYLEVK 137
**********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 79.846 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.5E-76 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.4E-50 | 22 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 6.5E-5 | 430 | 517 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.5E-4 | 431 | 504 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.71E-16 | 431 | 517 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.71E-16 | 618 | 724 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-17 | 623 | 725 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.08E-13 | 624 | 722 | No hit | No description |
| PROSITE profile | PS50297 | 17.263 | 630 | 724 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.3E-7 | 636 | 725 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.055 | 663 | 692 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.618 | 663 | 695 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 670 | 702 | 731 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.36 | 836 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.773 | 837 | 866 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0055 | 838 | 857 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.3E-7 | 838 | 887 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.0096 | 859 | 881 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.67 | 860 | 884 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.4E-4 | 862 | 881 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1026 aa Download sequence Send to blast |
MAEVRKYGLP NQPPDIPQIL LEAQNRWLRP TEICHILSNY KKFSIAPEPP NRPASGSLFL 60 FDRKILRYFR KDGHNWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRTYWL 120 LEEGFMNIVL VHYLEVKGKN NFSRVASKVS AVLKKLKRVQ DYLMLIVLHA QTLSLVRASV 180 SKEKFGATDN CRASSRYHPF VEMQQPVDGV MMNNMLGVSA PSAGYHGEMQ TTTANSDNHF 240 ATHYDIAGVF NEAGAGLRGV SKTLHDSVRF AEPYPEYSAE FMEPALYSSN ATMGSNNLDD 300 NSRLETFMSE ALYTNNLTQK EADALSAAGI MSSQAENNSY TDGIRYPLLK QSSLDLFKIE 360 PDGLKKFDSF SRWMSSELPE VADLDIKSSS DAFWSSTETV NVADGTSIPI NEQLDAFAVS 420 PSLSQDQLFS IIDVSPSYAC TGSRNKVLIT GTFLANKEHV ENCKWSCMFG DVEVPAEVLA 480 HGSLRCYTPV HLSGRVPFYV TCSNRVACSE VREFEFRDSD ARQMDTSDPQ TTGINEMHLH 540 IRLEKLLSLG PDDYEKYVMS DGKEKSEIIN TISSLMLDDK CLNQAVPLDE KEVSTARDQS 600 IEKLVKEKLY CWLVHKVHDE DKGPNVLGKE GQGVIHLVAA LGYDWAVRPI ITAGVKVNFR 660 DARGWTALHW AASCGRERTV GALIANGAES GLLTDPTPQF PAGRTAADLA SENGHKGIAG 720 FLAESALTSH LSALTLKESK DGNVKEICGL GGAEDFAESS SAQLAYRDSQ AESLKDSLSA 780 VRKSTQAAAR IFQAFRVESF HRKKVVEYGD DDCGLSDERT LSLVSIKNAK PGQNDGAHSA 840 AVRIQNKFRG WKGRKEFMII RQKIVKIQAH VRGHQVRKSY RKIVWSVGIV EKIILRWRRK 900 RRGLRGFQPV KQLEGPSPIQ QLEGPSQIQP AKEEEEDEYD YLKDGRKQAE GRLQRALARV 960 KSMTQYPEAR EQYSRIANRV TELQEPQAMM IQDDMQSDGA IADGGDFMAE LEELCGDEDA 1020 PMPTIL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
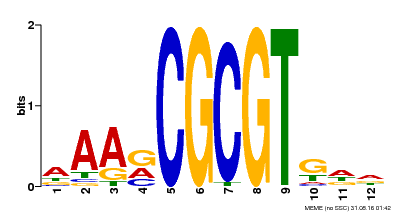
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN540903.1_FGP004 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK108461 | 0.0 | AK108461.1 Oryza sativa Japonica Group cDNA clone:002-143-D08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015647012.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | Q7XI44 | 0.0 | Q7XI44_ORYSJ; Putative anther ethylene-upregulated protein ER1 | ||||
| STRING | ORGLA07G0175700.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.3 | 0.0 | ethylene induced calmodulin binding protein | ||||



