 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN539461.1_FGP012 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 1196aa MW: 133867 Da PI: 6.6071 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.2 | 1.3e-31 | 269 | 323 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+rlrWt eLHerFveav++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYRl
KN539461.1_FGP012 269 KARLRWTLELHERFVEAVNKLDGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 323
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.393 | 266 | 326 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.7E-30 | 267 | 324 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.51E-16 | 267 | 323 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.6E-24 | 269 | 324 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.2E-7 | 271 | 322 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.1E-21 | 359 | 404 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Pfam | PF04937 | 1.1E-45 | 725 | 870 | IPR007021 | Domain of unknown function DUF659 |
| SuperFamily | SSF53098 | 6.98E-27 | 739 | 1120 | IPR012337 | Ribonuclease H-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1196 aa Download sequence Send to blast |
MSSQSVVAVK QIAAPDKIVE TCPSTKHSVH KLFDVKPDFQ GLMDDNLSSS SQSSSIKIEL 60 IRSSSLPNIL PFQKRSSEPE PESPLSHVSH PNVSEPVYSN SSTFCASLFS SSSMETEPCR 120 QLGTLPFLPH PPKCEQQVSA GHSSCSSLLV PGGDGDIGNA HDEPEQSDDL KDFLNLSGGD 180 ASDGSFHGEN NAMAFAEQME FQFLSEQLGI AITDNEESPR LDDIYGTPPQ LSSLPVSSCS 240 NQSVQKAGSP VKVQLSSPRS SSGSATTNKA RLRWTLELHE RFVEAVNKLD GPEKATPKGV 300 LKLMKVEGLT IYHVKSHLQK YRLAKYLPET KEDKKASSED KKSQSGSSGN DSVKKKNLQV 360 AEALRMQMEV QKQLHEQLEV QRQLQLRIEE HARYLQRILE EQHKVSISSN SLSLKPPAES 420 QPESPKPTSE KKEAESEAGA ATSAQPSSED KSPDAECKSS PPVCTTQDTQ RRRLEKEKHG 480 KTVDKETQKV SCNYCGKVVR TYNRLEHHLA GIRGNVYPCS LVPDSVRQSI KSSLEARKKD 540 WIARKIGKHK SSELPPVRNL SLTSAHACRP LLQPITSDTD REGSVSGHRC FVHFTNNLVQ 600 PGETTQVNAH DVSCNASSYQ QGGQTIEVTR PPYQNPCMMN KQPEMSSGQR IDPLSFSMEN 660 SSSQMQDSEF SKEPTNDYLN SQARKSIGKL IFEAGLEPGI LHLPSFKNMV DVLAWAQVSI 720 PTYESIMEEQ LREIQCHARD LKKHWEMNGC SVILDTWESR CGKSFISVLV HCSKGMLFIK 780 SMDVSDIIDD VDELAAMLFR VVEEVGVLNI VQIITNDESP YMQAAEHAVL KRYGYSFFFT 840 LCADHCINLL LENIAALDHV NEVLIKAREI TRFIYSHAVP MELKGKYIQG GEILSSSNLK 900 FVAMFITLGK LVSERINLVE MFSSPEWASS DLASRSSFRH VYEVVKTDNA FWSAAADILK 960 LTDPLITVLY KLEADNCPIG ILYDAMDCAK EDIKCNLRDK HGDYWPMVDE IWDHYLHTPV 1020 HAAGYILNPR IFYTDRFSYD TEIKSGTNAC VTRLAKNHYD PKKVAIQMDR YRRKSAPFDS 1080 DSAIQQTVEI PQVRWWSVHG TDTPELQTFA IRILSQTCFG ASTYNIDRSI SEQLHVVKRT 1140 YPEQERFRTM EYLHYNLRLA HCEPCVRGAS GAQQHSRLTS QLGDWISSGQ TTSYYK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
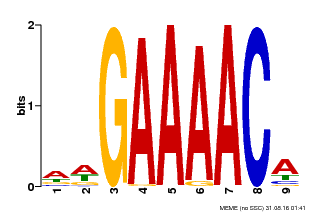
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN539461.1_FGP012 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012614 | 0.0 | CP012614.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 6 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015644149.1 | 0.0 | uncharacterized protein LOC4341991 isoform X1 | ||||
| TrEMBL | B8B263 | 0.0 | B8B263_ORYSI; Uncharacterized protein | ||||
| STRING | ORGLA06G0222700.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 7e-64 | G2-like family protein | ||||



