 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN539326.1_FGP004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 185aa MW: 20442.4 Da PI: 9.8936 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 99.7 | 1.9e-31 | 106 | 164 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-ST.T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsa.gCpvkkkversaedpkvveitYegeHnhe 59
D ++WrKYGqK++kgs+fpr+YY+C++ gCp++k+ver+ +dp+++++tYegeH+h+
KN539326.1_FGP004 106 ADDFSWRKYGQKPIKGSPFPRGYYKCSTLrGCPARKHVERDPTDPSMLIVTYEGEHRHT 164
699***********************9988****************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10533 | 1.5E-9 | 84 | 103 | IPR018872 | Zn-cluster domain |
| Gene3D | G3DSA:2.20.25.80 | 1.5E-32 | 93 | 164 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 29.792 | 100 | 166 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 3.01E-26 | 103 | 164 | IPR003657 | WRKY domain |
| SMART | SM00774 | 7.4E-37 | 105 | 165 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.7E-26 | 107 | 163 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:1990841 | Molecular Function | promoter-specific chromatin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 185 aa Download sequence Send to blast |
MDLMSGYGRV DEQVAIQEAA AAGLRGMEHL ILQLSQTGTS ERSPAPAPAQ EQQQQQVDCR 60 EITDMTVSKF KKVISMLNRT GHARKHRVKR TIRVPAISSK VADIPADDFS WRKYGQKPIK 120 GSPFPRGYYK CSTLRGCPAR KHVERDPTDP SMLIVTYEGE HRHTPSAAGQ DHPPAPPPPL 180 ALPLA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 2e-21 | 93 | 163 | 2 | 71 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts, when in complex with WRKY71, specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Represses specifically gibberellic acid (GA)-induced promoters in aleurone cells, probably by interfering with GAM1. {ECO:0000269|PubMed:16623886}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00464 | DAP | Transfer from AT4G31550 | Download |
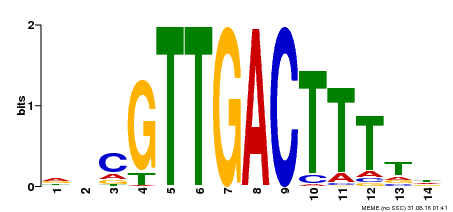
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN539326.1_FGP004 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA) in aleurone cells (PubMed:15618416, PubMed:16623886). Accumulates in response to uniconazole, a gibberellic acid (GA) biosynthesis inhibitor (PubMed:16623886). Repressed by GA (PubMed:15618416). {ECO:0000269|PubMed:15618416, ECO:0000269|PubMed:16623886}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK100954 | 1e-167 | AK100954.1 Oryza sativa Japonica Group cDNA clone:J023139N16, full insert sequence. | |||
| GenBank | AK121494 | 1e-167 | AK121494.1 Oryza sativa Japonica Group cDNA clone:J033000L17, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015633572.1 | 6e-58 | WRKY transcription factor WRKY51 | ||||
| Swissprot | Q0JEE2 | 5e-59 | WRK51_ORYSJ; WRKY transcription factor WRKY51 | ||||
| TrEMBL | A0A0E0DC00 | 4e-61 | A0A0E0DC00_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI04G19410.1 | 1e-69 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP617 | 38 | 171 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G31550.1 | 1e-47 | WRKY DNA-binding protein 11 | ||||



