 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN538863.1_FGP003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 163aa MW: 18741.4 Da PI: 9.1371 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 31.1 | 3.9e-10 | 97 | 130 | 22 | 55 |
SSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 22 nrypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++e++ +LA+++gL+ +q+ +WF N+R ++
KN538863.1_FGP003 97 WPYPTEEDKLRLAARTGLDPKQINNWFINQRKRH 130
69*****************************985 PP
| |||||||
| 2 | ELK | 34.3 | 5.1e-12 | 50 | 71 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsg+L+ L++EF+
KN538863.1_FGP003 50 ELKEMLLKKYSGCLSRLRSEFL 71
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03789 | 6.0E-9 | 50 | 71 | IPR005539 | ELK domain |
| SMART | SM01188 | 3.6E-6 | 50 | 71 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 10.423 | 50 | 70 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.843 | 70 | 133 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-13 | 72 | 137 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.2E-19 | 72 | 135 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-27 | 75 | 133 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.95E-13 | 82 | 134 | No hit | No description |
| Pfam | PF05920 | 2.4E-17 | 90 | 129 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 108 | 131 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 163 aa Download sequence Send to blast |
MSCKMVIPRS LVYNFMADEM VGSSDEDQCS GETDMLDIGQ EQSSRLADHE LKEMLLKKYS 60 GCLSRLRSEF LKKRKKGKLP KDARSALLEW WNTHYRWPYP TEEDKLRLAA RTGLDPKQIN 120 NWFINQRKRH WKPSDGMRFA LMEGVAGGSS GTTLYFDTGT IGP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 65 | 74 | LRSEFLKKRK |
| 2 | 71 | 75 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
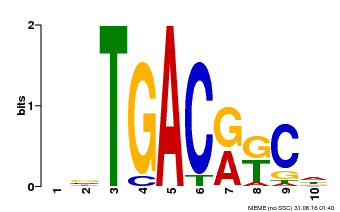
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN538863.1_FGP003 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241312 | 0.0 | AK241312.1 Oryza sativa Japonica Group cDNA, clone: J065141L09, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015634297.1 | 1e-105 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 1e-106 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | A0A0D3EMI9 | 1e-104 | A0A0D3EMI9_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0D9Y7A2 | 1e-104 | A0A0D9Y7A2_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0E0MV67 | 1e-104 | A0A0E0MV67_ORYRU; Uncharacterized protein | ||||
| TrEMBL | I1NM88 | 1e-104 | I1NM88_ORYGL; Uncharacterized protein | ||||
| STRING | OGLUM01G14210.1 | 1e-105 | (Oryza glumipatula) | ||||
| STRING | ORUFI01G13730.1 | 1e-105 | (Oryza rufipogon) | ||||
| STRING | OS01T0302500-01 | 1e-105 | (Oryza sativa) | ||||
| STRING | ORGLA01G0097800.1 | 1e-105 | (Oryza glaberrima) | ||||
| STRING | OBART01G11630.1 | 1e-105 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 9e-41 | KNOTTED-like from Arabidopsis thaliana | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



