 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OGLUM04G28480.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 1036aa MW: 114229 Da PI: 6.8279 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.7 | 4.7e-23 | 224 | 325 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k lt sd++++g +++p++ ae+ ++++++ ++l+++d++ + W++++iyr++++r++lt+GW+ Fv a++Lk+gD+v+F +++
OGLUM04G28480.1 224 FCKNLTASDTSTHGGFSVPRRAAEKLfpqldySMQPPN-QELIVRDLHDNMWTFRHIYRGQPKRHLLTTGWSLFVGAKRLKAGDSVLFI--RDE 314
89*********************999*****9555555.49************************************************..456 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
+ +l+++v+r+
OGLUM04G28480.1 315 KSQLLLGVRRA 325
77789999997 PP
| |||||||
| 2 | Auxin_resp | 117.3 | 1.3e-38 | 350 | 432 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++s F+++YnPr+s+s Fv++v++++ka++++ svGmRf m+fete+ss+rr++Gtvvg+sd+dp+rWpnSkWr+L+
OGLUM04G28480.1 350 AAHAASSGSSFTIYYNPRTSPSPFVIPVARYNKATYMQPSVGMRFAMMFETEESSKRRYTGTVVGISDYDPMRWPNSKWRNLQ 432
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.92E-45 | 214 | 353 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 4.8E-40 | 217 | 338 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.04E-20 | 223 | 324 | No hit | No description |
| SMART | SM01019 | 1.3E-22 | 224 | 326 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.3E-20 | 224 | 325 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.689 | 224 | 326 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.3E-34 | 350 | 432 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.2E-10 | 886 | 1020 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 8.5E-9 | 930 | 1012 | No hit | No description |
| PROSITE profile | PS51745 | 25.501 | 933 | 1017 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1036 aa Download sequence Send to blast |
MRNHGFPPRK VGIFRAKQPR SAASHTGAAA GERLGSQDQE REGKRRKGEE GSTTTLLRII 60 AIYCPHPTTT PTPVYSTPQP LLLPPREKAK TGVLRNAAAL LDEMQLMGET QGAKKVINSE 120 LWHACAGPLV CLPQRGSLVY YFPQGHSEQV AATTRKIPNS RIPNYPNLPS QLLCQVHNIT 180 LHADKDTDEV YAQMTLQPVN SETDVFPIPT LGAYTKSKHP TEYFCKNLTA SDTSTHGGFS 240 VPRRAAEKLF PQLDYSMQPP NQELIVRDLH DNMWTFRHIY RGQPKRHLLT TGWSLFVGAK 300 RLKAGDSVLF IRDEKSQLLL GVRRATRQQT MLSSSVLSTD SMHIGVLAAA AHAASSGSSF 360 TIYYNPRTSP SPFVIPVARY NKATYMQPSV GMRFAMMFET EESSKRRYTG TVVGISDYDP 420 MRWPNSKWRN LQVEWDEHGY GERPERVSIW DIETPENTLV FPSSTLNSKR QCLPGYGVSV 480 PGMEIGSANM SSFPRAQGNP YGSLQHIPAV GSELAIMLLN QSGQTLGSPL SFHQSSYSSI 540 IQNVKQNYIP PLTVSTSACL TKQESLPSDD AQHQFHMANM QNGDLEGSEV QPVIDLISES 600 KLNATSRDPR NTDSYTSRST SEQNSKGEPR GKTRRSKKGL PHKTVSEKSD LSSAPSWICD 660 NQQVGLESKL VGCDEQVNCG NIEDSSGALT QGNFVGQPHG HQVEQKGVLS PPKVESSKSP 720 DGGKSVNSFP NQGCFSQFID GLDWMTQPSY YQDSNVIQPA GVSENIFSSS ADIPPSMIAD 780 TMETFQASCL SDCLPNSIQE FISSPDLNSL TFLSPDMQNL EVQLQHDGSN LPSTSNSFVQ 840 MSFSEESASQ SENLSGLHME STHRSINTTS CSQPMSTGGF DAGMYSKLPR LKESQILSLP 900 EIHTNSMGTS ACSMDATEYS LDRSAKPMKP PVRTYTKVQK QGSVGRSIDV TGFRNYHELR 960 SAIACMFGLQ GKLEHPGSSE WKLVYVDYEN DVLLVGDDPW EEFINCVRCI RILSPSEVQQ 1020 MSENGMHVLN DCIQAA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 87 | 455 | 10 | 390 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
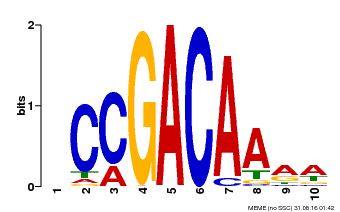
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB071292 | 0.0 | AB071292.1 Oryza sativa OsMP mRNA for Arabidopsis Monopteros-like protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025881052.1 | 0.0 | auxin response factor 11-like isoform X1 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A0A0D9ZS17 | 0.0 | A0A0D9ZS17_9ORYZ; Auxin response factor | ||||
| STRING | OGLUM04G28480.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



