 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB08G27190.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 996aa MW: 109972 Da PI: 7.1041 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130 | 8.7e-41 | 56 | 133 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C+adl++ak+yhrrhkvCe+h ka+++lv +++qrfCqqCsrfh lsefDe+krsCrrrLa+hn+rrrk+q+
OB08G27190.1 56 MCQVDDCRADLTNAKDYHRRHKVCEMHGKASKALVGNQMQRFCQQCSRFHPLSEFDEGKRSCRRRLAGHNRRRRKTQP 133
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-34 | 49 | 118 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.163 | 54 | 131 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 9.29E-38 | 55 | 134 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-30 | 57 | 130 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 996 aa Download sequence Send to blast |
MVVSPAATMS SSPSPPASSA PAQEPVVRPS KRVRSGSPGS ASGGGGGGNS GGSYPMCQVD 60 DCRADLTNAK DYHRRHKVCE MHGKASKALV GNQMQRFCQQ CSRFHPLSEF DEGKRSCRRR 120 LAGHNRRRRK TQPTDVASQL LLPGNQENAA NRTQDIVNLI TVIARLQGGN VGKLPSIPPI 180 PDKDNLVQII SKINSINNVN SASKSPPSEA VDLNATQGQH QDSVQRTTNG FEKQTNGFDK 240 QTVPSTMDLL TVLSTALATP NPDSNTSQSQ GSSDSSDNNK SKSHSTEPAN VVSSHEKSIR 300 VFSATRTNGI LESPPEVYKQ PEQETRPYLS LRLFGSTEED VPCKMDTANK YLSSESSNPL 360 DERSPSSSPP ITHKFFPIRS VHEEDRIADY GEDTATVEVS TSRAWHAPPL ELFKDSERPI 420 ENGSPPNPAY QSCYTSTSCS DHSPSTSNSD GQDRTGRIIF KLFGKEPSTI PGNLRGEIVN 480 WLKHSPTEME GYIRPGCLVL SIYLSMPTIA WDELQENLLQ RVNTLVQGSD LDFWRKGRFL 540 VRTDTQLVSY KDGTTRLSKS WRTWNTPELT FVSPIAVVGG RKTSLILKGR NLTIPGTQIH 600 CTNTGKYISK EVLCSAYPGT IYDDSGVETF DLPGEPHLVL GRYFIEVENR FRGNSFPVII 660 ANSSVCQELR SLEAELEGSQ FVDGSSDDQA HDARQLKPKD EVMHFLNELG WLFQKVAASA 720 SDGKSDPSVL DVIYFSTARF RYLLLFSSER DWCSLTRTLL EILVKRSLAS DELSQETLDM 780 LSEIHLLNRA VKRKSSQMAR LLVQFVVLCP DDSKLYPFLP NVAGPGGLTP LHLAASMEDA 840 EDIVDALTDD PQQVGLSCWH SVLDDDGQSP ETYAKLRNNN SYNELVAQKL VDRKNHQVTI 900 MVGKEEIHMD QPGNVGEKNK SAIQALQIRS CNQCAILDSG LLRRPLHSRG LLARPYIHSM 960 LAIAAVCVCV CVFMRALLRF NSGRTFKWER LDFGTI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-33 | 45 | 130 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00664 | PBM | Transfer from Potri.002G002400 | Download |
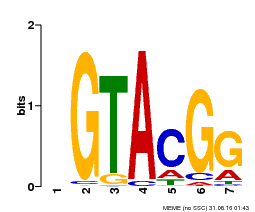
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB08G27190.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK242070 | 0.0 | AK242070.1 Oryza sativa Japonica Group cDNA, clone: J075133L10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006659584.2 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 15, partial | ||||
| Swissprot | A2YX04 | 0.0 | SPL15_ORYSI; Squamosa promoter-binding-like protein 15 | ||||
| Swissprot | Q6Z8M8 | 0.0 | SPL15_ORYSJ; Squamosa promoter-binding-like protein 15 | ||||
| TrEMBL | J3MUD8 | 0.0 | J3MUD8_ORYBR; Uncharacterized protein | ||||
| STRING | OB08G27190.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5456 | 38 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G20980.1 | 0.0 | squamosa promoter binding protein-like 14 | ||||



