 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB08G13700.1 | ||||||||
| Common Name | LOC102717637 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 718aa MW: 78916.2 Da PI: 6.137 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.8 | 3.2e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
OB08G13700.1 24 RERWTEAEHKRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.84E-16 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.147 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.6E-16 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.5E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.8E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.5E-12 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.03E-8 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 718 aa Download sequence Send to blast |
MEINSSGEEA VVKVRKPYTI TKQRERWTEA EHKRFLEALK LYGRAWQRIE EHVGTKTAVQ 60 IRSHAQKFFT KLEKEAINNG TSPGQAHDID IPPPRPKRKP NSPYPRKSSL SCETSIREVP 120 NDKATKSNLR SNSTSQMAGD AALEKLQRKE ISEKGSCSEV INLFEIPSAS FSSVNKGSSN 180 HDASRGMEPT KTEIKDVVIL ERDSIHNGVV KNAKDINDQE MERLNEMHIS SKSDHSHENC 240 LDASKQQSKP KPNSVEATYA DWSARRDSHY QMDRNGVTGI PATGTEGSHP DQTSDQMGGA 300 SGTMNQCIHP TLPVDPKFDR SDTAQPFPHN YTAFAPIMQC HCDQDAYRSF VNMSSTFSSM 360 LVSTLLSNPA IHAAARLAAS YWPAIDSNSP DPNQENISES AQGSHAGSPP NMASIVAATI 420 AAASAWWATQ GLLPLFPPPI AFPFVPAPSA PFPTADVQRA PEKDIDCPID NAQKELQETR 480 KQDKSEAMKV IVSSESDESG QGEVLLHTEL KISPADKDDT KTATGANTSE VFGNKKKQDR 540 SSCGSNTPSS SDIEADNAPE KQEKANDKAK QASCSNSSAG DTNHRRFRSS GSTSDSWKEV 600 SEEGRLAFDA LFSRERLPQS FSPPQAEGSK EVTKEDEDEV TTVTVDLNKN VTIIDHELDT 660 TDEPRASFPN ELSNLKLKSR RTGFKPYKRC SVEAKENRVP ASDEVGTKRI RLESETST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
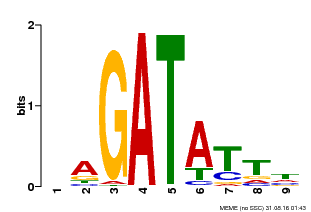
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB08G13700.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK242771 | 0.0 | AK242771.1 Oryza sativa Japonica Group cDNA, clone: J090053J23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006659145.1 | 0.0 | PREDICTED: protein CCA1 isoform X2 | ||||
| Refseq | XP_015695904.1 | 0.0 | PREDICTED: protein CCA1 isoform X2 | ||||
| TrEMBL | J3MQJ1 | 0.0 | J3MQJ1_ORYBR; Uncharacterized protein | ||||
| STRING | OB08G13700.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 1e-40 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 102717637 |



