 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB03G16740.1 | ||||||||
| Common Name | LOC102719565 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1047aa MW: 117094 Da PI: 5.8715 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.5 | 3.5e-55 | 19 | 135 | 2 | 117 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
lke ++rwl+++ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een +fqrr
OB03G16740.1 19 LKEaQHRWLRPTEICEILKNYRNFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENINFQRR 115
45559******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylev 117
+yw+Lee++ +ivlvhyle
OB03G16740.1 116 SYWMLEEDYMHIVLVHYLEI 135
*****************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.195 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-75 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.5E-48 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.6E-5 | 462 | 547 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 0.00764 | 462 | 549 | No hit | No description |
| Pfam | PF01833 | 1.1E-5 | 463 | 540 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 3.8E-15 | 463 | 548 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.34E-16 | 654 | 755 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.7E-18 | 654 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.71E-13 | 654 | 752 | No hit | No description |
| PROSITE profile | PS50088 | 8.79 | 660 | 692 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 18.086 | 660 | 764 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.013 | 693 | 722 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.843 | 693 | 725 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1400 | 732 | 761 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 4.51E-8 | 864 | 917 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.13 | 866 | 888 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.187 | 867 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0073 | 869 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 889 | 911 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 890 | 914 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 892 | 911 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1047 aa Download sequence Send to blast |
MAEGRRYAIA PQLDIEQILK EAQHRWLRPT EICEILKNYR NFRIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKSGSIDVLH CYYAHGEENI NFQRRSYWML 120 EEDYMHIVLV HYLEIKAGKL SSRSTGHDDI LQTSHVDSPL SQLPSQTTEG ESSVSGQASE 180 YDETESDIYS GGARHHPFSR TQQHENGGGS VIDHSIFSSY APASSLGSYQ GLQATAPNTG 240 FYSHGQDTLP VVLKESDLGT AFNGPNSQFD LSLWTEAMKP DIGTHQMPLY QPLVPPEQSP 300 FTEGSGIESF TFDEVYSNGL SIKDVGGDDT DGETPWQIPN ASVSFAAVDN FQQNDKSLEE 360 AISYPLLKTH SSGLSDILKE AINYPLLKTQ SSGLSDILKD GFKKNDSFTR WMSKELSEVD 420 DSQITSSSGV YWNSEEADNI IEASSSDQFT LAPVLAQDQL FSIVEFSPIW THAGSKTRVF 480 IKGKFLSSDE VKRFNWSCMF GEDEVPAEII ADDTLGCYSP SHKPGRVPFY VTCSNRLACS 540 EVREFEFRPQ YMDAPSPHGS TNKTYLQMRL DKLLSLEQDE IQSTLSNPTK EIVDLSKKIS 600 LLMMNNDDWS ELLKLADDNE PAIDDKQDQF LQKCIKEKLH IWLLHKVGDG SKGPSVLDEE 660 GQGVLHLAAA LGYDWAIRPT ITAGVNINFR DAHGWTALHW AAFCGRERTV VALIALGAAP 720 GAVTDPTPNF PSGSTPADLA SANGHKGISG FLAESSLTSH LQTLNLKEAM RSSAVEISGL 780 PAIVNVANRS TSPLAVEGLH TGSMGDSLGA FRNAAQAAAR IYQVFRMQSF QRKQAVQYED 840 DNGAISDERA MSLLSAKPSK PAQLDPLHVA ATRIQNKFRG WKGRKEFLLI RQRIVKIQAH 900 VRGHQVRKHY RKIVWSVGIV EKIILRWRRR GAGLRGFRPT ENADSTSSSS VDATQNKPAE 960 NDYDFLQEGR KQTEERLQKA LARVKSMVQY PEARDQYQRI LTVVTKMQES QALQEKMLEE 1020 STEMDEGLLM SEFQELLDDD MPTPGYF |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
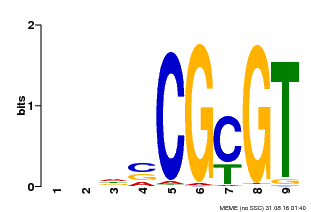
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB03G16740.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK066651 | 0.0 | AK066651.1 Oryza sativa Japonica Group cDNA clone:J013074G10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006649535.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 1-like | ||||
| TrEMBL | J3LKU8 | 0.0 | J3LKU8_ORYBR; Uncharacterized protein | ||||
| STRING | OB03G16740.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 102719565 |



