 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB02G21340.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 730aa MW: 85070.3 Da PI: 6.9887 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 49.2 | 1.7e-15 | 86 | 169 | 2 | 90 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHela 90
fY++ A+++GF v ++ skk ++g++ t+ Cs++gk++ ++++ + +++++tr +C k++ ++ d+k+++++l l+HnH+++
OB02G21340.1 86 FYKSCARKIGFGVIRRGSKKI-EDGKVRYFTLACSRQGKAQYTSTN---KFKPNPSTRQQCPTKVNFYLH-DEKFCISTLTLDHNHAVS 169
9*************9999988.8899****************9888...99******************9.9**************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.1E-12 | 86 | 170 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 9.1E-36 | 289 | 382 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.495 | 571 | 609 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.2E-4 | 583 | 603 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 0.0094 | 584 | 611 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 730 aa Download sequence Send to blast |
MSYHIFDENV VHTIFSYNYR TTIDGVGRNG DTNLCFDDEQ DHVISDDIFA VEEVESDSSA 60 LVADKTKLLE SAKGMLFDSE DSATCFYKSC ARKIGFGVIR RGSKKIEDGK VRYFTLACSR 120 QGKAQYTSTN KFKPNPSTRQ QCPTKVNFYL HDEKFCISTL TLDHNHAVSP SARHLRCHKE 180 LDLQAKRRLE LNDQAGIRIN KDFSSLVMQA GGYENLQFGQ KDCRNYLQDV QKLKLGAGDA 240 HAVYQYFLCM QSKDPNFFFV MDMDEDSRLK HVLWVDARSS ATYESFSDVV TFDTTYLTNK 300 YHMPFAPVGV NHHGESVLLG CALLSNEETE TFVWLFRSWL SCMSNKAPNA IITDQCRAMQ 360 NAIMEVFPEA RHRWCLWHIM KKIPEKLRGY LEYEDINSTL SNVVYDSLNR DDFDRGWMKM 420 IDEFSLQDNE WVVGLYDNRD LWVPTYVKDT FWVGMSSSQR SESVNAFFDG YVNARTTLKQ 480 FLEQCDNALR DKVEKENKSD CKSFQEAIPC ITHYEFEGQF QVAYTNKKYK EFQDELRGKI 540 YWYATLRKTE GSVHTFSVRE DRKIGEQRIV SKLLVLFNQE DCDLHCECRR FEFRGILCRH 600 IISILPLVEI EKVSSKYVIQ RWRKDFKHKH TFIKCSYDDQ LDTPIVRRFD TLCKRFNEVA 660 ENGSVSDALY NIVMDGLNEL QIKIDAHHQQ EIQQYQQNEK NKDMVPKQGK VVLSPISVRR 720 RGCPPSMRKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
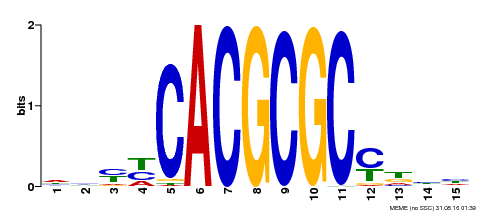
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB02G21340.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC136998 | 0.0 | AC136998.2 Oryza sativa (japonica cultivar-group) chromosome 11 BAC clone OSJNBa0082P17, complete sequence. | |||
| GenBank | AP014967 | 0.0 | AP014967.1 Oryza sativa Japonica Group DNA, chromosome 11, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024314597.1 | 0.0 | protein FAR1-RELATED SEQUENCE 6-like | ||||
| TrEMBL | J3LBW7 | 0.0 | J3LBW7_ORYBR; Uncharacterized protein | ||||
| STRING | OB02G21340.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP18 | 32 | 967 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 1e-124 | FAR1 family protein | ||||



