 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB02G10980.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 598aa MW: 68763.5 Da PI: 7.6022 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.4 | 2.5e-05 | 289 | 311 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ Cg sF + +Lk+H+ +H
OB02G10980.1 289 HNCQECGASFQKPAHLKQHMLSH 311
67*******************99 PP
| |||||||
| 2 | zf-C2H2 | 20.8 | 1e-06 | 317 | 341 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s++rk++L+rH+ +H
OB02G10980.1 317 FMCPleDCPFSYKRKDHLNRHLLKH 341
79*********************99 PP
| |||||||
| 3 | zf-C2H2 | 17.1 | 1.6e-05 | 346 | 371 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp Cg++F+ k n++rH + H
OB02G10980.1 346 FTCPmdGCGRTFNIKANMQRHVKEiH 371
89********************9988 PP
| |||||||
| 4 | zf-C2H2 | 14.1 | 0.00014 | 383 | 407 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ C+k+F s Lk+H +H
OB02G10980.1 383 FVCKeeGCNKVFMYASKLKKHEESH 407
89*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 17.3 | 1.3e-05 | 474 | 498 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ Cg sFs+ksnL++H++ H
OB02G10980.1 474 KCSfeGCGCSFSNKSNLTKHLKAcH 498
68999*****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF13041 | 5.6E-11 | 2 | 50 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 11.553 | 3 | 37 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 1.8E-5 | 5 | 38 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 4.5E-5 | 6 | 169 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 9.438 | 38 | 68 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 3.4E-4 | 40 | 67 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.774 | 74 | 104 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.027 | 77 | 101 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 5.47 | 106 | 136 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.87 | 140 | 174 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.35 | 149 | 171 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 32 | 243 | 265 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-5 | 286 | 308 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.466 | 289 | 316 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0078 | 289 | 311 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 291 | 311 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.15E-10 | 301 | 343 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-11 | 309 | 343 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0011 | 317 | 341 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.85 | 317 | 346 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 319 | 341 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.99E-6 | 345 | 372 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-6 | 346 | 372 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.0E-4 | 346 | 371 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.219 | 346 | 376 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 348 | 371 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.3E-7 | 381 | 407 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.138 | 383 | 407 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0068 | 383 | 407 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 385 | 407 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.1 | 415 | 440 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 417 | 440 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 13 | 443 | 464 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 9.29E-8 | 454 | 511 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.4E-7 | 455 | 495 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.967 | 473 | 503 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0033 | 473 | 498 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 475 | 498 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.5 | 504 | 530 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 506 | 530 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 598 aa Download sequence Send to blast |
MKDAFTWTTM ISSFTVQGDG KKAVELFWDM LRSGTPPNSV TFVSVLSACS HAGLIQEGRE 60 LFREMREVYH IDPRLEHYGC MVDLLGRGGL LEEAEALIDH MDVEPDIVIW RSLLSACLAR 120 GNDRLAEIAG KEIIRREPGD DGVYVLLWNM YASSNRWKEA LDMRKQMLSK KIYKTPGCSW 180 IEVDGVVHEF LVEDKTHDSR REIYGTLENM ARHLKMDPMY SGDEIDGDTK VEGATHYRDI 240 RRYKCEFCMV VRSKKCLIRA HMAAHHKEEL DESEIYKSNG EKIVQEGDHN CQECGASFQK 300 PAHLKQHMLS HLDERSFMCP LEDCPFSYKR KDHLNRHLLK HQGRLFTCPM DGCGRTFNIK 360 ANMQRHVKEI HEDGNATKSN QQFVCKEEGC NKVFMYASKL KKHEESHVKL DYVEVVCCEP 420 GCMKMFTNVE CLRAHNQACH QYIQCVICGE KYLKKNIKRH LRAHEGAPST ERIKCSFEGC 480 GCSFSNKSNL TKHLKACHDQ IKPFACRYTG CEKVFPYKHV RDNHEKSSAH VYTQANFMEM 540 DEQLRSCPRG GRKKKSVTVE TFTRKRVTMD GDASSLDNGT EYLRWLLSGG DDDASHAH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 3e-17 | 293 | 498 | 19 | 188 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 3e-17 | 293 | 498 | 19 | 188 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
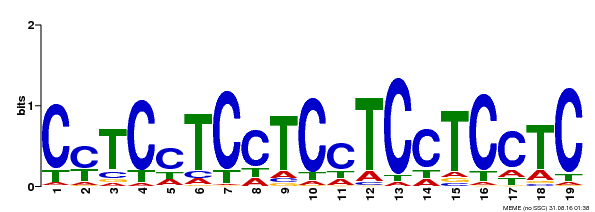
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB02G10980.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK100473 | 0.0 | AK100473.1 Oryza sativa Japonica Group cDNA clone:J023093M20, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006646782.1 | 0.0 | PREDICTED: transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 1e-114 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | J3L8Y1 | 0.0 | J3L8Y1_ORYBR; Uncharacterized protein | ||||
| STRING | OB02G10980.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-117 | transcription factor IIIA | ||||



