 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB01G12600.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 324aa MW: 36279.8 Da PI: 9.0573 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 41 | 4.6e-13 | 51 | 99 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ r r++lg+f +e Aa+a++ a+++++g
OB01G12600.1 51 SRYKGVVPQP-NGRWGAQIYE------RhSRVWLGTFPDEEAAARAYDVAALRFRG 99
78****9888.8*********......44*************************98 PP
| |||||||
| 2 | B3 | 90.9 | 9.6e-29 | 163 | 259 | 1 | 85 |
EEEE-..-HHHHTT-EE--HHH.HTT............---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh............ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+ tpsdv+k++rlv+pk+ ae+h + +++ l +ed +g++W+++++y+++s++yvltkGW++Fv+++gL+ gD+v F+ +
OB01G12600.1 163 FEKAVTPSDVGKLNRLVVPKQQAEKHfpfplrrssdaaAAATGKGVLLNFEDGDGKVWRFRYSYWNSSQSYVLTKGWSRFVREKGLRPGDTVAFSQS 259
89******************************9998885566799*************************************************954 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 8.5E-16 | 51 | 105 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 4.6E-8 | 51 | 99 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.47E-13 | 51 | 105 | No hit | No description |
| PROSITE profile | PS51032 | 20.508 | 52 | 107 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.7E-25 | 52 | 113 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.2E-19 | 52 | 105 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.9E-5 | 53 | 64 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.9E-5 | 73 | 89 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 2.8E-38 | 159 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 5.23E-29 | 160 | 274 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.39E-27 | 161 | 256 | No hit | No description |
| SMART | SM01019 | 4.0E-20 | 163 | 272 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.5E-26 | 163 | 267 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.915 | 163 | 273 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
MEQEGMVVSC NSGGSSSTES KPEDLVAMEE DEFQDLGGGH QAVELRVRPS SRYKGVVPQP 60 NGRWGAQIYE RHSRVWLGTF PDEEAAARAY DVAALRFRGR EAVTNRVTPE GASAGVLAFL 120 AAHSKAEIVD MLRKHTYDDE LQQGLRRGAR AQPTPRWARE PLFEKAVTPS DVGKLNRLVV 180 PKQQAEKHFP FPLRRSSDAA AAATGKGVLL NFEDGDGKVW RFRYSYWNSS QSYVLTKGWS 240 RFVREKGLRP GDTVAFSQSA WGTDKHLLID CKKMQRNQET HAIADDEARV VKLFGVDIAG 300 DKGVPQRICD SQKLQTVWDS FWQK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 7e-47 | 160 | 272 | 11 | 115 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
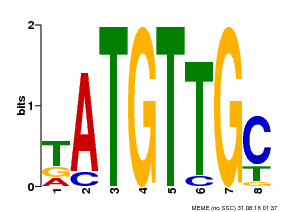
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB01G12600.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC211606 | 0.0 | AC211606.1 Oryza glaberrima clone OG_BBa0048H17, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006645486.1 | 0.0 | PREDICTED: putative AP2/ERF and B3 domain-containing protein Os01g0140700 | ||||
| Swissprot | Q9AWS7 | 1e-155 | Y1407_ORYSJ; Putative AP2/ERF and B3 domain-containing protein Os01g0140700 | ||||
| TrEMBL | J3KWA4 | 0.0 | J3KWA4_ORYBR; Uncharacterized protein | ||||
| STRING | OB01G12600.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 9e-95 | related to ABI3/VP1 2 | ||||



