 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART12G13720.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 350aa MW: 38010.2 Da PI: 7.0417 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.9 | 3e-15 | 36 | 83 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+lv + + G g+W+ +ar g+ R++k+c++rw +yl
OBART12G13720.1 36 KGLWSPEEDEKLVAYMLRSGQGSWSDVARNAGLQRCGKSCRLRWINYL 83
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.7 | 4e-16 | 89 | 132 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg+++++E++l+v +++ lG++ W+ Ia++++ gRt++++k++w++
OBART12G13720.1 89 RGAFSPQEEDLIVSLHAILGNR-WSQIAARLP-GRTDNEIKNFWNS 132
89********************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.314 | 31 | 83 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.72E-28 | 34 | 130 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.7E-11 | 35 | 85 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.3E-13 | 36 | 83 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-22 | 37 | 90 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.60E-9 | 39 | 83 | No hit | No description |
| PROSITE profile | PS51294 | 24.867 | 84 | 138 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.4E-15 | 88 | 136 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.1E-15 | 89 | 132 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.77E-11 | 91 | 131 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-25 | 91 | 138 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MRKPDCGGGG GAAKGGGALG VAGGNNAAVV GGKVRKGLWS PEEDEKLVAY MLRSGQGSWS 60 DVARNAGLQR CGKSCRLRWI NYLRPDLKRG AFSPQEEDLI VSLHAILGNR WSQIAARLPG 120 RTDNEIKNFW NSTIKKRLKI SSSSASPATT TDCASPPEHK LGALLPLPDQ VCGVDTSPPP 180 PFFHDHSISI KQAYYGSTGA HHHHHAIAAM DGSSLIGDHH HHSSSILFGG ASVPPLLDHQ 240 TILNDDDDHP NKTGSNTTAA TLSSNITDNS NSNKNNSDNN NNISSSCCIS LMNSSSNMIY 300 WEGHHQQQQQ QHQMLLQQQQ HMSRNVMGEW DLEELMKDVS SLPFLDFQVE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-27 | 36 | 138 | 7 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00599 | PBM | Transfer from AT5G12870 | Download |
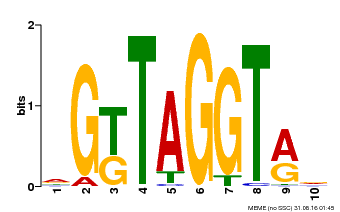
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AL713900 | 0.0 | AL713900.4 Oryza sativa chromosome 12, . BAC OJ1126_F07 of library Monsanto from chromosome 12 of cultivar Nipponbare of ssp. japonica of Oryza sativa (rice), complete sequence. | |||
| GenBank | AP014968 | 0.0 | AP014968.1 Oryza sativa Japonica Group DNA, chromosome 12, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012620 | 0.0 | CP012620.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 12 sequence. | |||
| GenBank | JN634084 | 0.0 | JN634084.1 Oryza sativa Japonica Group secondary wall MYB46 transcription factor mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015619488.1 | 1e-146 | transcription factor MYB83 | ||||
| TrEMBL | A0A0D3HV16 | 0.0 | A0A0D3HV16_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART12G13720.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9821 | 35 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G12870.1 | 1e-64 | myb domain protein 46 | ||||



