 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART04G21010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 546aa MW: 56961.1 Da PI: 6.2012 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 15.3 | 5.8e-05 | 118 | 135 | 2 | 19 |
EETTTTEEESSHHHHHHH CS
zf-C2H2 2 kCpdCgksFsrksnLkrH 19
+C++Cgk F++++++ H
OBART04G21010.1 118 VCKNCGKEFTSWEHFLEH 135
7**************999 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 0.96 | 29 | 51 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.94E-6 | 29 | 51 | No hit | No description |
| PROSITE profile | PS50157 | 9.536 | 29 | 51 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF13912 | 0.0045 | 30 | 51 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 31 | 51 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.94E-6 | 112 | 136 | No hit | No description |
| SMART | SM00355 | 140 | 117 | 137 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.141 | 117 | 144 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.71E-8 | 257 | 280 | No hit | No description |
| SMART | SM00355 | 0.13 | 258 | 280 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.242 | 258 | 285 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF13912 | 3.0E-9 | 258 | 282 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 260 | 280 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.71E-8 | 366 | 393 | No hit | No description |
| SMART | SM00355 | 3.7 | 371 | 393 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF13912 | 4.6E-11 | 371 | 394 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.182 | 371 | 393 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 373 | 393 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 546 aa Download sequence Send to blast |
MHHHGQMQPS PSSPRAPPPT AAQASGYKHF CRVCNKGFTC GSALGGHMRA HGVGDGDGLG 60 ADDDDDDDDD SLGDEAVRRA RGGADDPWNA GGPSSSGAAT HVYELRTNPN RVTRSRQVCK 120 NCGKEFTSWE HFLEHGKCSS GEDDADGEED PAPAAGWLKG KRSRRCKGTG VDLSPTPSAC 180 AAGEEEDLAN CLVLLSSSKV DQAGVTEAEQ PSSSSASKEH KRLITFMEPT TYVLDTVMAL 240 PPPAPAPQYV STVPRGMFEC KACKKVFSSH QALGGHRASH KKVKGCFAAK LESNAAEVAE 300 PSHHAEVADR SEDNPAKATS DARRNVHASM DGDGNAGTSD AAAELSMAIV PIEPPVAALA 360 AAPLKKKGKM HECSVCHRLF TSGQALGGHK RCHWLTSSSA DHTASVPPLA DDLVPLSFRP 420 MLDAPEPALD LSIAANPPLL ASAATVRPKV GGSSFHLDAP PPVYIPSSPA IPSQRNKATA 480 TTGSQNANDA VGLSTAAAED EADSTTVKRA RLSDLKDVSM AGETTPWLQV GIGSSSRGGA 540 DDNDKE |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 363 | 369 | LKKKGKM |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00621 | PBM | Transfer from AT2G17180 | Download |
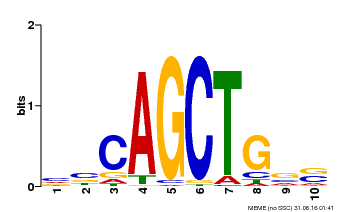
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012612 | 0.0 | CP012612.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 4 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015634601.1 | 0.0 | uncharacterized protein LOC4336604 | ||||
| TrEMBL | A0A0D3FYQ1 | 0.0 | A0A0D3FYQ1_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART04G21010.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3524 | 32 | 62 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G17180.1 | 7e-21 | C2H2-like zinc finger protein | ||||



