 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART03G05550.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 314aa MW: 33448.4 Da PI: 7.721 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63 | 6.6e-20 | 27 | 76 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+WvAeIr+p +kr+r +lg+f taeeAa a+++a+++l+g
OBART03G05550.1 27 EYRGVRQRT-WGKWVAEIREP---NKRTRLWLGSFATAEEAALAYDEAARRLYG 76
69****999.**********9...337*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.38E-26 | 26 | 86 | No hit | No description |
| PROSITE profile | PS51032 | 22.55 | 27 | 84 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 6.8E-14 | 27 | 76 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.8E-36 | 27 | 90 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.12E-21 | 27 | 85 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.0E-30 | 27 | 85 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.1E-10 | 28 | 39 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.1E-10 | 50 | 66 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 314 aa Download sequence Send to blast |
MESYGRKRAW KKGPTRGKGG PQNAACEYRG VRQRTWGKWV AEIREPNKRT RLWLGSFATA 60 EEAALAYDEA ARRLYGPDAF LNLPHLRAAS AAAAHQRLRW LPASAAAAAA GGRRRRPRLR 120 PPQPQRAAQR PRHTPEAAGA QELVVVADQA AAAYADARQP ASAASPHLVT LLHRHQQRRL 180 RGSASAHVAL EQAMAATAAM ESAPCDDDAA VVGFGADKPQ LDLKEFLQQI GVLKADDDGA 240 TGKNGAVHGD DGELADAFGF GGSGEFDWDA LAADMSDIAG GHGGALGANG GFQMDDLHEV 300 EQFGGCMPIP IWDI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-16 | 28 | 90 | 6 | 68 | ATERF1 |
| 3gcc_A | 2e-16 | 28 | 90 | 6 | 68 | ATERF1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 118 | GRRRRPRL |
| 2 | 113 | 120 | RRRPRLRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that binds to the DNA sequence 5'-[AG]CCGAC-3' of the cis-acting dehydration-responsive element (DRE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00411 | DAP | Transfer from AT3G57600 | Download |
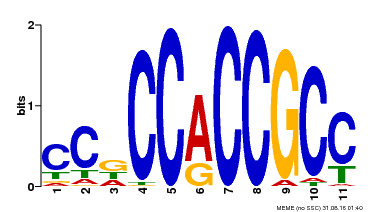
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not induced by high-salt and drought stresses. {ECO:0000269|PubMed:20049613}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC090484 | 0.0 | AC090484.4 Genomic sequence for Oryza sativa, Nipponbare strain, clone OSJNBa0061L19, from chromosome 3, complete sequence. | |||
| GenBank | AP014959 | 0.0 | AP014959.1 Oryza sativa Japonica Group DNA, chromosome 3, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025879844.1 | 1e-139 | dehydration-responsive element-binding protein 2E | ||||
| Swissprot | Q10R18 | 1e-140 | DRE2E_ORYSJ; Dehydration-responsive element-binding protein 2E | ||||
| TrEMBL | A0A0D3FEH5 | 0.0 | A0A0D3FEH5_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART03G05550.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9575 | 29 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G57600.1 | 6e-34 | ERF family protein | ||||



