 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009630328.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 423aa MW: 45877.5 Da PI: 6.7237 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 106.1 | 5.7e-33 | 29 | 150 | 3 | 89 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt.....................ssssaseceae 76
g+kdrhsk++T++g+RdRRvRlsa++a++f+d+qd+LG+d++sk+++WL+++a++ai+el ++ +
XP_009630328.1 29 GRKDRHSKVCTAKGPRDRRVRLSAHTAIQFYDVQDRLGYDRPSKAVDWLIKKAQAAIDELEELppwkpttgpianstnfdqekkK---------- 113
89*************************************************************9888876655433333222220.......... PP
TCP 77 sssssasnsssg...........................k 89
++++ +++ +
XP_009630328.1 114 -QKENFDTQ--QndvnvsiggssskramimlgnesearvL 150
.11111111..11233444444333444444433322220 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.2E-29 | 29 | 166 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 33.621 | 30 | 88 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009793 | Biological Process | embryo development ending in seed dormancy | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 423 aa Download sequence Send to blast |
MGTSSKFGLR NSGGEIVEVQ GGHIIRSIGR KDRHSKVCTA KGPRDRRVRL SAHTAIQFYD 60 VQDRLGYDRP SKAVDWLIKK AQAAIDELEE LPPWKPTTGP IANSTNFDQE KKKQKENFDT 120 QQNDVNVSIG GSSSKRAMIM LGNESEARVL HQTNSGNSSG DNANASFLPQ SLDSDTIADT 180 IKSFFPVGCS NESTSSSAMQ FQSFQEHNLL SRTSSHSQDL RLSLQSFQDP ILLQQQAQHH 240 NNPRNHTEEE QTHFQASNFF TGSTPFGFDA SGWSVSEHQQ QPVEQRPAAA RAGGDSTGIA 300 LCGGAVAGAA GGYLFNSPPA PPLLQQLFGQ NQFFSQRGPL QSSYSPSIRA WIDPSAIAIA 360 TADPIHHHQA MLPMYSASIS GIGFASELGG FSGFRIPARI QGEIEEHDGI SDKPSSASSD 420 SRH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 2e-20 | 35 | 88 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 2e-20 | 35 | 88 | 1 | 54 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor playing a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164) (PubMed:12931144, PubMed:17307931). Required during early steps of embryogenesis (PubMed:15634699). Participates in ovule develpment (PubMed:25378179). Activates LOX2 expression by binding to the 5'-GGACCA-3' motif found in its promoter (PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:15634699, ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:18816164, ECO:0000269|PubMed:25378179}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00062 | PBM | Transfer from AT3G15030 | Download |
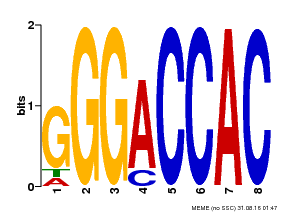
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by the miRNA miR-JAW/miR319 (PubMed:12931144, PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:18816164}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975451 | 6e-85 | HG975451.1 Solanum pennellii chromosome ch12, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009630327.1 | 0.0 | PREDICTED: transcription factor TCP4-like | ||||
| Refseq | XP_016463776.1 | 0.0 | PREDICTED: transcription factor TCP4-like | ||||
| Refseq | XP_016463777.1 | 0.0 | PREDICTED: transcription factor TCP4-like | ||||
| Swissprot | Q8LPR5 | 6e-98 | TCP4_ARATH; Transcription factor TCP4 | ||||
| TrEMBL | A0A1S3ZHP1 | 0.0 | A0A1S3ZHP1_TOBAC; transcription factor TCP4-like | ||||
| STRING | XP_009630327.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G15030.3 | 4e-85 | TCP family protein | ||||



